DISCOVR EPIC L2
The matchups in this notebook require xarray version > 2026.02
The DSCOVR Earth Polychromatic Imaging Camera (EPIC) observes the sunlit Earth from the Sun–Earth L1 point and captures full-disk images of the planet in 10 UV–visible–near-IR spectral bands roughly every 1–2 hours.
Level-2 products are geophysical retrievals derived from EPIC radiances.
Cloud properties: cloud mask, cloud optical thickness, cloud effective pressure/height/temperature, cloud phase
Aerosols: aerosol optical depth, aerosol index
Trace gases: total column ozone, sulfur dioxide (SO₂)
Surface/vegetation: surface reflectivity, UV aerosol index, vegetation indices (e.g., NDVI)
This notebook will use two DISCOVR EPIC Level 2 products.
DSCOVR_EPIC_L2_AER, Version 03, aerosols
DSCOVR_EPIC_L2_TO3 Version 03, total column ozone (TO₃)
These granules are big and the grid is not reqular. A KD tree is used for level 2 data and this can be slow.
Note: In a virtual machine in AWS us-west-2, where NASA cloud data is, the point matchups are faster. In Colab, say, your compute is not in the same data region nor provider, and the same matchups might take 10x longer.
Prerequisites
The examples here use NASA EarthData and you need to have an account with EarthData. Make sure you can login.
# if needed
! pip install point - collocation cartopy "xarray>=2026.2" -- quiet
import earthaccess
earthaccess . login ()
<earthaccess.auth.Auth at 0x7fefa734ad20>
Check the xarray version
import xarray as xr
xr . __version__
DSCOVR_EPIC_L2_AER
Create some points
Random global and in 2024.
import pandas as pd
url = (
"https://raw.githubusercontent.com/"
"fish-pace/point-collocation/main/"
"examples/fixtures/points_1000.csv"
)
df_points = pd . read_csv (
url ,
parse_dates = [ "time" ]
)
df = df_points [
( df_points [ "time" ] . dt . year == 2024 )
]
print ( len ( df ))
df . head ()
lat
lon
time
land
35
9.100711
11.216734
2024-07-21
True
43
-67.209878
-62.734423
2024-04-16
False
83
87.228338
-29.718344
2024-10-13
False
190
34.701655
-20.040258
2024-04-21
False
222
-8.043907
41.779801
2024-08-31
False
Create the plan
We look for granules within 1 hour of our points. There are 14 points that have matches and some have multiple matches (granules).
%% time
import point_collocation as pc
short_name = "DSCOVR_EPIC_L2_AER"
plan = pc . plan (
df ,
data_source = "earthaccess" ,
source_kwargs = {
"short_name" : short_name ,
"version" : "03"
},
time_buffer = "1h"
)
CPU times: user 1.55 s, sys: 136 ms, total: 1.68 s
Wall time: 7.36 s
Plan: 31 points → 17 unique granule(s)
Points with 0 matches : 20
Points with >1 matches: 6
Time buffer: 0 days 01:00:00
Look at the variables
We will open a file with datatree and see what groups it has.
ds = plan . open_dataset ( 1 , open_method = "datatree" )
ds
open_method: {'xarray_open': 'datatree', 'open_kwargs': {'chunks': {}, 'engine': 'h5netcdf', 'decode_timedelta': False}, 'merge': None, 'coords': 'auto', 'set_coords': True, 'dim_renames': None, 'auto_align_phony_dims': None}
<xarray.DataTree>
Group: /
├── Group: /HDFEOS
│ ├── Group: /HDFEOS/ADDITIONAL
│ │ └── Group: /HDFEOS/ADDITIONAL/FILE_ATTRIBUTES
│ │ Attributes: (12/22)
│ │ ACF Filename: acf_epic_sm_ffxt_aug2019.txt
│ │ AIRSCO_filename1: AIRS.2016.05.08.L3.RetStd_IR001.v6.0.31.0.G161301816...
│ │ AIRSCO_filename2: AIRS.2016.05.07.L3.RetStd_IR001.v6.0.31.0.G161291747...
│ │ AuthorAffiliation: NASA/GSFC
│ │ AuthorName: Omar Torres
│ │ EarthSunDistance: 0.9862366
│ │ ... ...
│ │ ProcessingCenter: TLCF ...
│ │ ProcessingHost: Linux ominate.gsfc.nasa.gov 2.6.18-410.el5PAE #1 SMP...
│ │ ProductionDateTime: 2024-02-11T01:29:46.0Z
│ │ ShortName: EPICAERUV
│ │ Time_Band340nm: 2024-02-08 01:57:26
│ │ Time_Band388nm: 2024-02-08 01:56:59
│ └── Group: /HDFEOS/SWATHS
│ └── Group: /HDFEOS/SWATHS/Aerosol NearUV Swath
│ ├── Group: /HDFEOS/SWATHS/Aerosol NearUV Swath/Data Fields
│ │ Dimensions: (phony_dim_0: 2048, phony_dim_1: 2048,
│ │ phony_dim_2: 5, phony_dim_3: 3,
│ │ phony_dim_4: 2)
│ │ Dimensions without coordinates: phony_dim_0, phony_dim_1, phony_dim_2,
│ │ phony_dim_3, phony_dim_4
│ │ Data variables: (12/29)
│ │ AIRSCO_Flags (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ │ AIRSL3COvalue (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ │ AerosolAbsOpticalDepthVsHeight (phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3) float32 252MB dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
│ │ AerosolCorrCloudOpticalDepth (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ │ AerosolOpticalDepthOverCloud (phony_dim_0, phony_dim_1, phony_dim_3) float32 50MB dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
│ │ AerosolOpticalDepthVsHeight (phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3) float32 252MB dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
│ │ ... ...
│ │ Reflectivity (phony_dim_0, phony_dim_1, phony_dim_4) float32 34MB dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
│ │ Residue (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ │ SurfaceAlbedo (phony_dim_0, phony_dim_1, phony_dim_4) float32 34MB dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
│ │ SurfaceType (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ │ UVAerosolIndex (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ │ Wavelength (phony_dim_3) float32 12B dask.array<chunksize=(3,), meta=np.ndarray>
│ └── Group: /HDFEOS/SWATHS/Aerosol NearUV Swath/Geolocation Fields
│ Dimensions: (phony_dim_5: 2048, phony_dim_6: 2048)
│ Dimensions without coordinates: phony_dim_5, phony_dim_6
│ Data variables:
│ Latitude (phony_dim_5, phony_dim_6) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ Longitude (phony_dim_5, phony_dim_6) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ RelativeAzimuthAngle (phony_dim_5, phony_dim_6) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ SnowIce_fraction (phony_dim_5, phony_dim_6) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ SolarZenithAngle (phony_dim_5, phony_dim_6) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ TerrainPressure (phony_dim_5, phony_dim_6) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
│ ViewingZenithAngle (phony_dim_5, phony_dim_6) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
└── Group: /HDFEOS INFORMATION
Dimensions: ()
Data variables:
StructMetadata.0 |S32000 32kB ...
Attributes:
HDFEOSVersion: HDFEOS_5.1.15 Groups: (2)
Groups: (2)
Groups: (1)
Groups: (2)
Dimensions: phony_dim_0 : 2048phony_dim_1 : 2048phony_dim_2 : 5phony_dim_3 : 3phony_dim_4 : 2
Data variables: (29)
AIRSCO_Flags
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 255 Title : AIRSCO_Flags Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
AIRSL3COvalue
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -9999.0 Title : AIRS CO L3 data Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
AerosolAbsOpticalDepthVsHeight
(phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Aerosol Absorption Optical Depth (tau_abs) at 5 levels Units : NoUnits Array Chunk Bytes 240.00 MiB 15.00 MiB Shape (2048, 2048, 5, 3) (256, 1024, 5, 3) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 1 3 5 2048
AerosolCorrCloudOpticalDepth
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Cloud Optical Depth Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
AerosolOpticalDepthOverCloud
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Aerosol Optical Depth Over Cloud Units : NoUnits Array Chunk Bytes 48.00 MiB 2.00 MiB Shape (2048, 2048, 3) (256, 1024, 2) Dask graph 32 chunks in 2 graph layers Data type float32 numpy.ndarray
3 2048 2048
AerosolOpticalDepthVsHeight
(phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Aerosol Optical Depth (tau) at 5 levels Units : NoUnits Array Chunk Bytes 240.00 MiB 15.00 MiB Shape (2048, 2048, 5, 3) (256, 1024, 5, 3) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 1 3 5 2048
AerosolSingleScattAlbVsHeight
(phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Aerosol Single Scattering Albedo (omega0) at 5 levels Units : NoUnits Array Chunk Bytes 240.00 MiB 15.00 MiB Shape (2048, 2048, 5, 3) (256, 1024, 5, 3) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 1 3 5 2048
AerosolType
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 255 Title : Aerosol Type Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
AlgorithmFlagsVsHeight
(phony_dim_0, phony_dim_1, phony_dim_2)
float32
dask.array<chunksize=(256, 1024, 3), meta=np.ndarray>
MissingValue : 65535 Title : Algorithm Flags Vs Height Units : NoUnits Array Chunk Bytes 80.00 MiB 3.00 MiB Shape (2048, 2048, 5) (256, 1024, 3) Dask graph 32 chunks in 2 graph layers Data type float32 numpy.ndarray
5 2048 2048
AlgorithmFlags_AerosolIndex
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 65535 Title : Algorithm Flags for AerosolIndex Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
CloudFraction
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Cloud Fraction Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
CloudOpticalDepth
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Cloud Optical Depth Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
FinalAerosolAbsOpticalDepth
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Best Aerosol Absorption Optical Depth (tau_abs) Units : NoUnits Array Chunk Bytes 48.00 MiB 3.00 MiB Shape (2048, 2048, 3) (256, 1024, 3) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
3 2048 2048
FinalAerosolLayerHeight
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Final Aerosol Layer Height (km) Units : km Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
FinalAerosolOpticalDepth
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Best Aerosol Optical Depth (tau) Units : NoUnits Array Chunk Bytes 48.00 MiB 3.00 MiB Shape (2048, 2048, 3) (256, 1024, 3) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
3 2048 2048
FinalAerosolSingleScattAlb
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Best Aerosol Single Scattering Albedo (omega0) Units : NoUnits Array Chunk Bytes 48.00 MiB 3.00 MiB Shape (2048, 2048, 3) (256, 1024, 3) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
3 2048 2048
FinalAlgorithmFlags
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 65535 Title : Final Algorithm Flags Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
FinalAlgorithmFlagsEPICAERAC
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 65535 Title : Final Algorithm Flags EPICAERAC Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
FinalImRefIdx
(phony_dim_0, phony_dim_1, phony_dim_4)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : FinalImRefIdx Units : NoUnits Array Chunk Bytes 32.00 MiB 2.00 MiB Shape (2048, 2048, 2) (256, 1024, 2) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2 2048 2048
HeightFlags
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 255 Title : Height Flags Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
ImaRefractiveIndex
(phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Imaginary refractive Index Units : NoUnits Array Chunk Bytes 240.00 MiB 15.00 MiB Shape (2048, 2048, 5, 3) (256, 1024, 5, 3) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 1 3 5 2048
InputSSAEPICAERAC
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Input SSA for EPICAERAC Units : NoUnits Array Chunk Bytes 48.00 MiB 2.00 MiB Shape (2048, 2048, 3) (256, 1024, 2) Dask graph 32 chunks in 2 graph layers Data type float32 numpy.ndarray
3 2048 2048
NormRadiance
(phony_dim_0, phony_dim_1, phony_dim_4)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Normalized Radiance Units : NoUnits Array Chunk Bytes 32.00 MiB 2.00 MiB Shape (2048, 2048, 2) (256, 1024, 2) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2 2048 2048
Reflectivity
(phony_dim_0, phony_dim_1, phony_dim_4)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Lambert Equivalent Reflectivity Units : NoUnits Array Chunk Bytes 32.00 MiB 2.00 MiB Shape (2048, 2048, 2) (256, 1024, 2) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2 2048 2048
Residue
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Residue Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
SurfaceAlbedo
(phony_dim_0, phony_dim_1, phony_dim_4)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Surface Albedo Units : NoUnits Array Chunk Bytes 32.00 MiB 2.00 MiB Shape (2048, 2048, 2) (256, 1024, 2) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2 2048 2048
SurfaceType
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 65535 Title : Surface Type Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
UVAerosolIndex
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : UV Aerosol Index Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
Wavelength
(phony_dim_3)
float32
dask.array<chunksize=(3,), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Wavelength Units : nm Array Chunk Bytes 12 B 12 B Shape (3,) (3,) Dask graph 1 chunks in 2 graph layers Data type float32 numpy.ndarray
3 1
Dimensions: phony_dim_5 : 2048phony_dim_6 : 2048
Data variables: (7)
Latitude
(phony_dim_5, phony_dim_6)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Geodetic Latitude (deg) Units : deg Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
Longitude
(phony_dim_5, phony_dim_6)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Geodetic Longitude (deg) Units : deg Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
RelativeAzimuthAngle
(phony_dim_5, phony_dim_6)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Relative (sun + 180 - view) Azimuth Angle (deg) Units : deg(EastofNorth) Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
SnowIce_fraction
(phony_dim_5, phony_dim_6)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : SnowIce_fraction) Units : NoUnits Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
SolarZenithAngle
(phony_dim_5, phony_dim_6)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Solar Zenith Angle (deg) Units : deg Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
TerrainPressure
(phony_dim_5, phony_dim_6)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Terrain Pressure (mbar) Units : mbar Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
ViewingZenithAngle
(phony_dim_5, phony_dim_6)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Viewing Zenith Angle (deg) Units : deg Array Chunk Bytes 16.00 MiB 1.00 MiB Shape (2048, 2048) (256, 1024) Dask graph 16 chunks in 2 graph layers Data type float32 numpy.ndarray
2048 2048
The data are in /HDFEOS/SWATHS/Aerosol NearUV Swath/Data Fields and lat/lon are in /HDFEOS/SWATHS/Aerosol NearUV Swath/Geolocation Fields. We create a profile. We open a dataset with the profile to make sure it looks good.
%% time
discovr_epic_aer = {
'xarray_open' : 'dataset' ,
'merge' : [ '/HDFEOS/SWATHS/Aerosol NearUV Swath/Geolocation Fields' , '/HDFEOS/SWATHS/Aerosol NearUV Swath/Data Fields' ],
'open_kwargs' : { 'phony_dims' : 'access' }
}
ds = plan . open_dataset ( 1 , open_method = discovr_epic_aer )
ds
open_method: {'xarray_open': 'dataset', 'merge': ['/HDFEOS/SWATHS/Aerosol NearUV Swath/Geolocation Fields', '/HDFEOS/SWATHS/Aerosol NearUV Swath/Data Fields'], 'open_kwargs': {'phony_dims': 'access', 'chunks': {}, 'engine': 'h5netcdf', 'decode_timedelta': False}, 'coords': 'auto', 'set_coords': True, 'dim_renames': None, 'auto_align_phony_dims': None, 'merge_kwargs': {}}
CPU times: user 803 ms, sys: 99.6 ms, total: 903 ms
Wall time: 4.35 s
<xarray.Dataset> Size: 2GB
Dimensions: (phony_dim_0: 2048, phony_dim_1: 2048,
phony_dim_2: 5, phony_dim_3: 3,
phony_dim_4: 2)
Coordinates:
Latitude (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
Longitude (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
Dimensions without coordinates: phony_dim_0, phony_dim_1, phony_dim_2,
phony_dim_3, phony_dim_4
Data variables: (12/34)
RelativeAzimuthAngle (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
SnowIce_fraction (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
SolarZenithAngle (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
TerrainPressure (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
ViewingZenithAngle (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
AIRSCO_Flags (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
... ...
Reflectivity (phony_dim_0, phony_dim_1, phony_dim_4) float32 34MB dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
Residue (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
SurfaceAlbedo (phony_dim_0, phony_dim_1, phony_dim_4) float32 34MB dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
SurfaceType (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
UVAerosolIndex (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 1024), meta=np.ndarray>
Wavelength (phony_dim_3) float32 12B dask.array<chunksize=(3,), meta=np.ndarray> Dimensions: phony_dim_0 : 2048phony_dim_1 : 2048phony_dim_2 : 5phony_dim_3 : 3phony_dim_4 : 2Coordinates: (2)
Latitude
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Geodetic Latitude (deg) Units : deg
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
Longitude
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Geodetic Longitude (deg) Units : deg
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
Data variables: (34)
RelativeAzimuthAngle
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Relative (sun + 180 - view) Azimuth Angle (deg) Units : deg(EastofNorth)
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
SnowIce_fraction
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : SnowIce_fraction) Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
SolarZenithAngle
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Solar Zenith Angle (deg) Units : deg
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
TerrainPressure
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Terrain Pressure (mbar) Units : mbar
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
ViewingZenithAngle
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Viewing Zenith Angle (deg) Units : deg
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
AIRSCO_Flags
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 255 Title : AIRSCO_Flags Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
AIRSL3COvalue
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -9999.0 Title : AIRS CO L3 data Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
AerosolAbsOpticalDepthVsHeight
(phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Aerosol Absorption Optical Depth (tau_abs) at 5 levels Units : NoUnits
Array
Chunk
Bytes
240.00 MiB
15.00 MiB
Shape
(2048, 2048, 5, 3)
(256, 1024, 5, 3)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
1
3
5
2048
AerosolCorrCloudOpticalDepth
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Cloud Optical Depth Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
AerosolOpticalDepthOverCloud
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Aerosol Optical Depth Over Cloud Units : NoUnits
Array
Chunk
Bytes
48.00 MiB
2.00 MiB
Shape
(2048, 2048, 3)
(256, 1024, 2)
Dask graph
32 chunks in 2 graph layers
Data type
float32 numpy.ndarray
3
2048
2048
AerosolOpticalDepthVsHeight
(phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Aerosol Optical Depth (tau) at 5 levels Units : NoUnits
Array
Chunk
Bytes
240.00 MiB
15.00 MiB
Shape
(2048, 2048, 5, 3)
(256, 1024, 5, 3)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
1
3
5
2048
AerosolSingleScattAlbVsHeight
(phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Aerosol Single Scattering Albedo (omega0) at 5 levels Units : NoUnits
Array
Chunk
Bytes
240.00 MiB
15.00 MiB
Shape
(2048, 2048, 5, 3)
(256, 1024, 5, 3)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
1
3
5
2048
AerosolType
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 255 Title : Aerosol Type Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
AlgorithmFlagsVsHeight
(phony_dim_0, phony_dim_1, phony_dim_2)
float32
dask.array<chunksize=(256, 1024, 3), meta=np.ndarray>
MissingValue : 65535 Title : Algorithm Flags Vs Height Units : NoUnits
Array
Chunk
Bytes
80.00 MiB
3.00 MiB
Shape
(2048, 2048, 5)
(256, 1024, 3)
Dask graph
32 chunks in 2 graph layers
Data type
float32 numpy.ndarray
5
2048
2048
AlgorithmFlags_AerosolIndex
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 65535 Title : Algorithm Flags for AerosolIndex Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
CloudFraction
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Cloud Fraction Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
CloudOpticalDepth
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Cloud Optical Depth Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
FinalAerosolAbsOpticalDepth
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Best Aerosol Absorption Optical Depth (tau_abs) Units : NoUnits
Array
Chunk
Bytes
48.00 MiB
3.00 MiB
Shape
(2048, 2048, 3)
(256, 1024, 3)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
3
2048
2048
FinalAerosolLayerHeight
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Final Aerosol Layer Height (km) Units : km
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
FinalAerosolOpticalDepth
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Best Aerosol Optical Depth (tau) Units : NoUnits
Array
Chunk
Bytes
48.00 MiB
3.00 MiB
Shape
(2048, 2048, 3)
(256, 1024, 3)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
3
2048
2048
FinalAerosolSingleScattAlb
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Best Aerosol Single Scattering Albedo (omega0) Units : NoUnits
Array
Chunk
Bytes
48.00 MiB
3.00 MiB
Shape
(2048, 2048, 3)
(256, 1024, 3)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
3
2048
2048
FinalAlgorithmFlags
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 65535 Title : Final Algorithm Flags Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
FinalAlgorithmFlagsEPICAERAC
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 65535 Title : Final Algorithm Flags EPICAERAC Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
FinalImRefIdx
(phony_dim_0, phony_dim_1, phony_dim_4)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : FinalImRefIdx Units : NoUnits
Array
Chunk
Bytes
32.00 MiB
2.00 MiB
Shape
(2048, 2048, 2)
(256, 1024, 2)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2
2048
2048
HeightFlags
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 255 Title : Height Flags Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
ImaRefractiveIndex
(phony_dim_0, phony_dim_1, phony_dim_2, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 5, 3), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Imaginary refractive Index Units : NoUnits
Array
Chunk
Bytes
240.00 MiB
15.00 MiB
Shape
(2048, 2048, 5, 3)
(256, 1024, 5, 3)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
1
3
5
2048
InputSSAEPICAERAC
(phony_dim_0, phony_dim_1, phony_dim_3)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Input SSA for EPICAERAC Units : NoUnits
Array
Chunk
Bytes
48.00 MiB
2.00 MiB
Shape
(2048, 2048, 3)
(256, 1024, 2)
Dask graph
32 chunks in 2 graph layers
Data type
float32 numpy.ndarray
3
2048
2048
NormRadiance
(phony_dim_0, phony_dim_1, phony_dim_4)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Normalized Radiance Units : NoUnits
Array
Chunk
Bytes
32.00 MiB
2.00 MiB
Shape
(2048, 2048, 2)
(256, 1024, 2)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2
2048
2048
Reflectivity
(phony_dim_0, phony_dim_1, phony_dim_4)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Lambert Equivalent Reflectivity Units : NoUnits
Array
Chunk
Bytes
32.00 MiB
2.00 MiB
Shape
(2048, 2048, 2)
(256, 1024, 2)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2
2048
2048
Residue
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Residue Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
SurfaceAlbedo
(phony_dim_0, phony_dim_1, phony_dim_4)
float32
dask.array<chunksize=(256, 1024, 2), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Surface Albedo Units : NoUnits
Array
Chunk
Bytes
32.00 MiB
2.00 MiB
Shape
(2048, 2048, 2)
(256, 1024, 2)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2
2048
2048
SurfaceType
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : 65535 Title : Surface Type Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
UVAerosolIndex
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 1024), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : UV Aerosol Index Units : NoUnits
Array
Chunk
Bytes
16.00 MiB
1.00 MiB
Shape
(2048, 2048)
(256, 1024)
Dask graph
16 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
Wavelength
(phony_dim_3)
float32
dask.array<chunksize=(3,), meta=np.ndarray>
MissingValue : -1.2676506e+30 Title : Wavelength Units : nm
Array
Chunk
Bytes
12 B
12 B
Shape
(3,)
(3,)
Dask graph
1 chunks in 2 graph layers
Data type
float32 numpy.ndarray
3
1
Let's get the matchups
%% time
res = pc . matchup ( plan [ 0 : 10 ],
variables = [ "UVAerosolIndex" ],
open_method = discovr_epic_aer )
CPU times: user 49.5 s, sys: 497 ms, total: 50 s
Wall time: 1min 11s
# full res shows more info about matchups, like the granule lat lon
var = 'UVAerosolIndex'
res [[ 'lat' , 'lon' , 'time' , 'pc_id' , var ]] . dropna ( subset = [ var ]) . head ()
lat
lon
time
pc_id
UVAerosolIndex
5
6.631233
-138.494233
2024-05-08
245
1.105056
6
-20.064168
-173.807753
2024-02-16
273
0.151248
8
34.247526
-166.904639
2024-08-21
342
0.469726
9
34.247526
-166.904639
2024-08-21
342
0.536712
This takes about 25 seconds per point/granule. We will want to use parallel processing to speed this up so we can make many granule I/O requests at once.
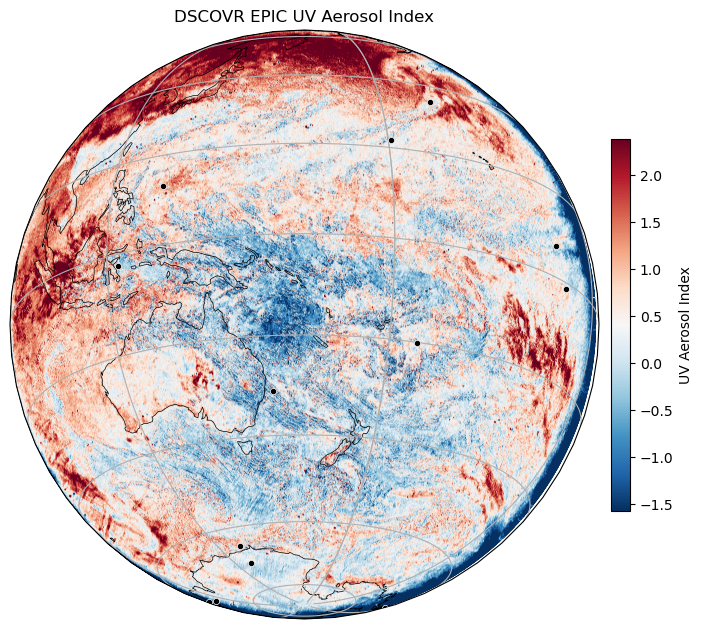
Let's make a plot
We could do ds.UVAerosolIndex.plot(robust=True) which is fast but also ugly plot. The following makes a prettier plot.
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER , LATITUDE_FORMATTER
import numpy as np
# mask fill values
fill = ds . UVAerosolIndex . attrs [ "MissingValue" ]
uvai = ds . UVAerosolIndex . where ( ds . UVAerosolIndex != fill )
lat = ds . Latitude . values
lon = ds . Longitude . values
val = uvai . values
vmin , vmax = np . nanpercentile ( val , [ 2 , 98 ])
# flatten for scatter
lat = lat . flatten ()
lon = lon . flatten ()
val = val . flatten ()
# projection center (mean location of valid pixels)
lat2d = ds . Latitude . values
lon2d = ds . Longitude . values
iy = lat2d . shape [ 0 ] // 2
ix = lat2d . shape [ 1 ] // 2
lat_m = float ( lat2d [ iy , ix ])
lon_m = float ( lon2d [ iy , ix ])
proj = ccrs . Orthographic ( central_longitude = lon_m , central_latitude = lat_m )
fig = plt . figure ( figsize = ( 8 , 8 ))
ax = plt . axes ( projection = proj )
# coastlines
ax . coastlines ( linewidth = 0.5 )
# gridlines
grid = ax . gridlines ( draw_labels = False )
grid . xformatter = LONGITUDE_FORMATTER
grid . yformatter = LATITUDE_FORMATTER
# plot swath
im = ax . scatter (
lon ,
lat ,
c = val ,
s = 1 ,
cmap = "RdBu_r" ,
vmin = vmin ,
vmax = vmax ,
transform = ccrs . PlateCarree ()
)
# points
ax . scatter (
df [ "lon" ],
df [ "lat" ],
color = "black" ,
s = 20 ,
edgecolor = "white" ,
linewidth = 0.5 ,
transform = ccrs . PlateCarree (),
zorder = 10
)
plt . colorbar ( im , fraction = 0.03 , pad = 0.02 , label = "UV Aerosol Index" )
plt . title ( "DSCOVR EPIC UV Aerosol Index" )
plt . show ()
DSCOVR_EPIC_L2_TO3
Total ozone.
%% time
import point_collocation as pc
short_name = "DSCOVR_EPIC_L2_TO3"
plan = pc . plan (
df ,
data_source = "earthaccess" ,
source_kwargs = {
"short_name" : short_name ,
"version" : "03"
},
time_buffer = "1h"
)
CPU times: user 816 ms, sys: 69.6 ms, total: 886 ms
Wall time: 11.9 s
Look at variables
All the data is in the base group so we do not need to pass in open_method; the default open_dataset will work fine. Our variable of interest is "Ozone".
ds = plan . open_dataset ( 1 , open_method = "dataset" )
ds
open_method: {'xarray_open': 'dataset', 'open_kwargs': {'chunks': {}, 'engine': 'h5netcdf', 'decode_timedelta': False}, 'merge': None, 'coords': 'auto', 'set_coords': True, 'dim_renames': None, 'auto_align_phony_dims': None}
<xarray.Dataset> Size: 549MB
Dimensions: (phony_dim_0: 2048, phony_dim_1: 2048,
phony_dim_2: 4, phony_dim_3: 11,
phony_dim_4: 26, phony_dim_5: 3,
phony_dim_6: 1, phony_dim_7: 8)
Coordinates:
Latitude (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 256), meta=np.ndarray>
Longitude (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 256), meta=np.ndarray>
Dimensions without coordinates: phony_dim_0, phony_dim_1, phony_dim_2,
phony_dim_3, phony_dim_4, phony_dim_5,
phony_dim_6, phony_dim_7
Data variables: (12/27)
AlgorithmFlag (phony_dim_0, phony_dim_1) int16 8MB dask.array<chunksize=(256, 256), meta=np.ndarray>
CalibrationCoef (phony_dim_2) float32 16B dask.array<chunksize=(4,), meta=np.ndarray>
CloudPressure (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 256), meta=np.ndarray>
ColumnWeightFunctionPercent (phony_dim_0, phony_dim_1, phony_dim_3) uint8 46MB dask.array<chunksize=(256, 256, 11), meta=np.ndarray>
ControlFileContents (phony_dim_4) object 208B dask.array<chunksize=(26,), meta=np.ndarray>
DNDomega (phony_dim_0, phony_dim_1, phony_dim_5) float32 50MB dask.array<chunksize=(256, 256, 3), meta=np.ndarray>
... ...
SlopeOfReflectivity (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 256), meta=np.ndarray>
SolarAzimuthAngle (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 256), meta=np.ndarray>
SolarZenithAngle (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 256), meta=np.ndarray>
TerrainPressure (phony_dim_0, phony_dim_1) float32 17MB dask.array<chunksize=(256, 256), meta=np.ndarray>
Wavelength (phony_dim_2) float32 16B dask.array<chunksize=(4,), meta=np.ndarray>
YearDaySeconds (phony_dim_5) int32 12B dask.array<chunksize=(3,), meta=np.ndarray> Dimensions: phony_dim_0 : 2048phony_dim_1 : 2048phony_dim_2 : 4phony_dim_3 : 11phony_dim_4 : 26phony_dim_5 : 3phony_dim_6 : 1phony_dim_7 : 8Coordinates: (2)
Latitude
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
Longitude
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
Data variables: (27)
AlgorithmFlag
(phony_dim_0, phony_dim_1)
int16
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
8.00 MiB
128.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
int16 numpy.ndarray
2048
2048
CalibrationCoef
(phony_dim_2)
float32
dask.array<chunksize=(4,), meta=np.ndarray>
Array
Chunk
Bytes
16 B
16 B
Shape
(4,)
(4,)
Dask graph
1 chunks in 2 graph layers
Data type
float32 numpy.ndarray
4
1
CloudPressure
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
ColumnWeightFunctionPercent
(phony_dim_0, phony_dim_1, phony_dim_3)
uint8
dask.array<chunksize=(256, 256, 11), meta=np.ndarray>
Array
Chunk
Bytes
44.00 MiB
704.00 kiB
Shape
(2048, 2048, 11)
(256, 256, 11)
Dask graph
64 chunks in 2 graph layers
Data type
uint8 numpy.ndarray
11
2048
2048
ControlFileContents
(phony_dim_4)
object
dask.array<chunksize=(26,), meta=np.ndarray>
Array
Chunk
Bytes
208 B
208 B
Shape
(26,)
(26,)
Dask graph
1 chunks in 2 graph layers
Data type
object numpy.ndarray
26
1
DNDomega
(phony_dim_0, phony_dim_1, phony_dim_5)
float32
dask.array<chunksize=(256, 256, 3), meta=np.ndarray>
Array
Chunk
Bytes
48.00 MiB
768.00 kiB
Shape
(2048, 2048, 3)
(256, 256, 3)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
3
2048
2048
DNDreflectivity
(phony_dim_0, phony_dim_1, phony_dim_2)
float32
dask.array<chunksize=(256, 256, 4), meta=np.ndarray>
Array
Chunk
Bytes
64.00 MiB
1.00 MiB
Shape
(2048, 2048, 4)
(256, 256, 4)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
4
2048
2048
ErrorFlag
(phony_dim_0, phony_dim_1)
int16
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
8.00 MiB
128.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
int16 numpy.ndarray
2048
2048
ErythemalUV
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
LERTerrPres
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
NValue
(phony_dim_0, phony_dim_1, phony_dim_2)
float32
dask.array<chunksize=(256, 256, 4), meta=np.ndarray>
Array
Chunk
Bytes
64.00 MiB
1.00 MiB
Shape
(2048, 2048, 4)
(256, 256, 4)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
4
2048
2048
NoABandCloud
(phony_dim_6)
int32
dask.array<chunksize=(1,), meta=np.ndarray>
Array
Chunk
Bytes
4 B
4 B
Shape
(1,)
(1,)
Dask graph
1 chunks in 2 graph layers
Data type
int32 numpy.ndarray
1
1
NvalueAdjust
(phony_dim_2)
float32
dask.array<chunksize=(4,), meta=np.ndarray>
Array
Chunk
Bytes
16 B
16 B
Shape
(4,)
(4,)
Dask graph
1 chunks in 2 graph layers
Data type
float32 numpy.ndarray
4
1
Ozone
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
OzoneStep1
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
RadiativeCloudFraction
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
RamanCorrection
(phony_dim_7, phony_dim_2)
float32
dask.array<chunksize=(8, 4), meta=np.ndarray>
Array
Chunk
Bytes
128 B
128 B
Shape
(8, 4)
(8, 4)
Dask graph
1 chunks in 2 graph layers
Data type
float32 numpy.ndarray
4
8
Reflectivity
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
Residual
(phony_dim_0, phony_dim_1, phony_dim_5)
float32
dask.array<chunksize=(256, 256, 3), meta=np.ndarray>
Array
Chunk
Bytes
48.00 MiB
768.00 kiB
Shape
(2048, 2048, 3)
(256, 256, 3)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
3
2048
2048
SatelliteAzimuthAngle
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
SatelliteZenithAngle
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
SlopeOfReflectivity
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
SolarAzimuthAngle
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
SolarZenithAngle
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
TerrainPressure
(phony_dim_0, phony_dim_1)
float32
dask.array<chunksize=(256, 256), meta=np.ndarray>
Array
Chunk
Bytes
16.00 MiB
256.00 kiB
Shape
(2048, 2048)
(256, 256)
Dask graph
64 chunks in 2 graph layers
Data type
float32 numpy.ndarray
2048
2048
Wavelength
(phony_dim_2)
float32
dask.array<chunksize=(4,), meta=np.ndarray>
Array
Chunk
Bytes
16 B
16 B
Shape
(4,)
(4,)
Dask graph
1 chunks in 2 graph layers
Data type
float32 numpy.ndarray
4
1
YearDaySeconds
(phony_dim_5)
int32
dask.array<chunksize=(3,), meta=np.ndarray>
Array
Chunk
Bytes
12 B
12 B
Shape
(3,)
(3,)
Dask graph
1 chunks in 2 graph layers
Data type
int32 numpy.ndarray
3
1
%% time
res = pc . matchup ( plan [ 0 : 10 ], variables = [ "Ozone" ])
CPU times: user 35.8 s, sys: 942 ms, total: 36.8 s
Wall time: 1min 34s
var = 'Ozone'
res [[ 'lat' , 'lon' , 'time' , 'pc_id' , var ]] . dropna ( subset = [ var ]) . head ()
lat
lon
time
pc_id
Ozone
5
6.631233
-138.494233
2024-05-08
245
249.927444
6
-20.064168
-173.807753
2024-02-16
273
233.636459
8
34.247526
-166.904639
2024-08-21
342
316.187988
9
34.247526
-166.904639
2024-08-21
342
314.785095
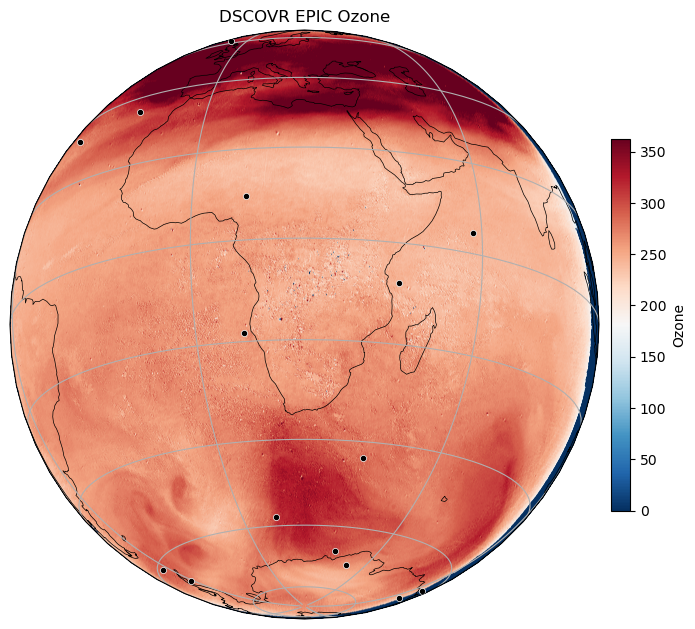
Make a plot
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER , LATITUDE_FORMATTER
import numpy as np
var = 'Ozone'
uvai = ds [ var ]
lat = ds . Latitude . values
lon = ds . Longitude . values
val = uvai . values
vmin , vmax = np . nanpercentile ( val , [ 2 , 98 ])
vmin = 0
# flatten for scatter
lat = lat . flatten ()
lon = lon . flatten ()
val = val . flatten ()
# projection center (mean location of valid pixels)
lat2d = ds . Latitude . values
lon2d = ds . Longitude . values
iy = lat2d . shape [ 0 ] // 2
ix = lat2d . shape [ 1 ] // 2
lat_m = float ( lat2d [ iy , ix ])
lon_m = float ( lon2d [ iy , ix ])
proj = ccrs . Orthographic ( central_longitude = lon_m , central_latitude = lat_m )
fig = plt . figure ( figsize = ( 8 , 8 ))
ax = plt . axes ( projection = proj )
# coastlines
ax . coastlines ( linewidth = 0.5 )
# gridlines
grid = ax . gridlines ( draw_labels = False )
grid . xformatter = LONGITUDE_FORMATTER
grid . yformatter = LATITUDE_FORMATTER
# plot swath
im = ax . scatter (
lon ,
lat ,
c = val ,
s = 1 ,
cmap = "RdBu_r" ,
vmin = vmin ,
vmax = vmax ,
transform = ccrs . PlateCarree ()
)
# points
ax . scatter (
df [ "lon" ],
df [ "lat" ],
color = "black" ,
s = 20 ,
edgecolor = "white" ,
linewidth = 0.5 ,
transform = ccrs . PlateCarree (),
zorder = 10
)
plt . colorbar ( im , fraction = 0.03 , pad = 0.02 , label = "Ozone" )
plt . title ( "DSCOVR EPIC Ozone" )
plt . show ()