NASA Earthdata Search and Discovery for PACE data#
Author: Eli Holmes (NOAA) Last updated: Jan 22, 2026
📘 Learning Objectives
How to authenticate with
earthaccess.login()How to use
earthaccess.search_data()to search for data using spatial and temporal filtersHow to explore and work with search results
Use
xarray.open_dataset()to load Level 3 dataUse xarray and datatree to load Level 2 data
(Optional) Plotting and managing memory.
Summary#
In this example we will use the earthaccess library to search for data collections from NASA Earthdata. earthaccess is a Python library that simplifies data discovery and access to NASA Earth science data by providing an abstraction layer for NASA’s Common Metadata Repository (CMR) API Search API. The library makes searching for data more approachable by using a simpler notation instead of low level HTTP queries. earthaccess takes the trouble out of Earthdata Login authentication, makes search easier, and provides a stream-line way to download or stream search results into an xarray object.
For more on earthaccess visit the earthaccess GitHub page and/or the earthaccess documentation site. Be aware that earthaccess is under active development.
Prerequisites#
An Earthdata Login account is required to access data from NASA Earthdata. Please visit https://urs.earthdata.nasa.gov to register and manage your Earthdata Login account. This account is free to create and only takes a moment to set up.
# If running in Colab, uncomment (delete the #) and run this line
# !pip install xarray earthaccess cartopy
Import Required Packages#
import earthaccess
import xarray as xr
Authenticate to NASA Earthdata#
This will ask for your username and password. persist=True will cause a .netrc file to be added to your home directory with your Earthdata login information.
auth = earthaccess.login(persist=True)
Search for data earthaccess.search_data()#
We want to search NASA Earthdata for collections using the OCI instrument. Use https://search.earthdata.nasa.gov/search?fi=OCI
There are multiple keywords we can use to discovery data from collections. We are going to focus on short_name here, but you can also use concept_id, and doi.
Search by sensor instrument="oci"#
results = earthaccess.search_datasets(instrument="oci")
for item in results[1:20]:
summary = item.summary()
print(summary["short-name"])
PACE_OCI_L2_AOP_NRT
PACE_OCI_L2_AER_UAA
PACE_OCI_L2_AOP
PACE_OCI_L2_BGC_NRT
PACE_OCI_L3M_CHL_NRT
PACE_OCI_L2_BGC
PACE_OCI_L2_CLOUD_MASK
PACE_OCI_L2_SFREFL_NRT
PACE_OCI_L3M_CHL
PACE_OCI_L1C_SCI
PACE_OCI_L2_CLOUD
PACE_OCI_L3M_RRS
PACE_OCI_L2_CLOUD_NRT
PACE_OCI_L2_IOP
PACE_OCI_L2_SFREFL
PACE_OCI_L2_UVAI_UAA_NRT
PACE_OCI_L3M_AER_UAA_NRT
PACE_OCI_L3M_CLOUD
PACE_OCI_L4M_MOANA
Search by short name short_name = ""#
Let’s look at Level 2 Apparent Optical Properties. Short name is PACE_OCI_L2_AOP. Notice L2 in the name. I can restrict to a time period using temporal with start and end date.
# Level 2 data
results = earthaccess.search_data(
short_name = 'PACE_OCI_L2_AOP',
temporal = ("2025-03-05", "2025-03-05")
)
len(results)
143
Why are there so many?? Level 2 data is swath data not a global grid. For swath data (level 2), we need to specify a bounding box so we only get the swath(s) over our region of interest. Here we specify a bounding box for the U.S. Great Lakes. Now we see there are 3 files for that day and box.
# Level 2 data
# bounding_box = (lon_min, lat_min, lon_max, lat_max)
results = earthaccess.search_data(
short_name = 'PACE_OCI_L2_AOP',
temporal = ("2025-03-05", "2025-03-05"),
bounding_box = (-90.0, 40.0, -75.0, 47.0)
)
len(results)
3
The Level 3 mapped data is global data on a grid (no swaths and no tiles) so there should be one file per day, right? Notice the L3M in the short_name.
# Level 3 data
results = earthaccess.search_data(
short_name = 'PACE_OCI_L3M_RRS',
temporal = ("2025-03-05", "2025-03-05")
)
len(results)
16
Why so many? There are month, 8-day, day and 2 resolutions. Look at the url for Data: in the results object.
results[0:5]
[Collection: {'Version': '3.1', 'ShortName': 'PACE_OCI_L3M_RRS'}
Spatial coverage: {'HorizontalSpatialDomain': {'Geometry': {'BoundingRectangles': [{'EastBoundingCoordinate': 180, 'SouthBoundingCoordinate': -90, 'NorthBoundingCoordinate': 90, 'WestBoundingCoordinate': -180}]}}}
Temporal coverage: {'RangeDateTime': {'BeginningDateTime': '2024-12-21T00:00:00Z', 'EndingDateTime': '2025-03-20T23:59:59Z'}}
Size(MB): 3143.3699989318848
Data: ['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20241221_20250320.L3m.SNWI.RRS.V3_1.Rrs.4km.nc'],
Collection: {'ShortName': 'PACE_OCI_L3M_RRS', 'Version': '3.1'}
Spatial coverage: {'HorizontalSpatialDomain': {'Geometry': {'BoundingRectangles': [{'NorthBoundingCoordinate': 90, 'SouthBoundingCoordinate': -90, 'WestBoundingCoordinate': -180, 'EastBoundingCoordinate': 180}]}}}
Temporal coverage: {'RangeDateTime': {'BeginningDateTime': '2024-12-21T00:00:00Z', 'EndingDateTime': '2025-03-20T23:59:59Z'}}
Size(MB): 579.0568437576294
Data: ['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20241221_20250320.L3m.SNWI.RRS.V3_1.Rrs.0p1deg.nc'],
Collection: {'ShortName': 'PACE_OCI_L3M_RRS', 'Version': '3.1'}
Spatial coverage: {'HorizontalSpatialDomain': {'Geometry': {'BoundingRectangles': [{'NorthBoundingCoordinate': 90, 'SouthBoundingCoordinate': -90, 'WestBoundingCoordinate': -180, 'EastBoundingCoordinate': 180}]}}}
Temporal coverage: {'RangeDateTime': {'BeginningDateTime': '2025-02-02T00:00:00Z', 'EndingDateTime': '2025-03-05T23:59:59Z'}}
Size(MB): 594.6580295562744
Data: ['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250202_20250305.L3m.R32.RRS.V3_1.Rrs.0p1deg.nc'],
Collection: {'Version': '3.1', 'ShortName': 'PACE_OCI_L3M_RRS'}
Spatial coverage: {'HorizontalSpatialDomain': {'Geometry': {'BoundingRectangles': [{'NorthBoundingCoordinate': 90, 'WestBoundingCoordinate': -180, 'EastBoundingCoordinate': 180, 'SouthBoundingCoordinate': -90}]}}}
Temporal coverage: {'RangeDateTime': {'BeginningDateTime': '2025-02-02T00:00:00Z', 'EndingDateTime': '2025-03-05T23:59:59Z'}}
Size(MB): 3193.7952098846436
Data: ['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250202_20250305.L3m.R32.RRS.V3_1.Rrs.4km.nc'],
Collection: {'ShortName': 'PACE_OCI_L3M_RRS', 'Version': '3.1'}
Spatial coverage: {'HorizontalSpatialDomain': {'Geometry': {'BoundingRectangles': [{'NorthBoundingCoordinate': 90, 'SouthBoundingCoordinate': -90, 'WestBoundingCoordinate': -180, 'EastBoundingCoordinate': 180}]}}}
Temporal coverage: {'RangeDateTime': {'BeginningDateTime': '2025-02-10T00:00:00Z', 'EndingDateTime': '2025-03-13T23:59:59Z'}}
Size(MB): 598.9511699676514
Data: ['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250210_20250313.L3m.R32.RRS.V3_1.Rrs.0p1deg.nc']]
# here is a way to just get the urls
[res.data_links() for res in results]
[['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20241221_20250320.L3m.SNWI.RRS.V3_1.Rrs.4km.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20241221_20250320.L3m.SNWI.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250202_20250305.L3m.R32.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250202_20250305.L3m.R32.RRS.V3_1.Rrs.4km.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250210_20250313.L3m.R32.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250210_20250313.L3m.R32.RRS.V3_1.Rrs.4km.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250218_20250321.L3m.R32.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250218_20250321.L3m.R32.RRS.V3_1.Rrs.4km.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250226_20250305.L3m.8D.RRS.V3_1.Rrs.4km.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250226_20250305.L3m.8D.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250226_20250329.L3m.R32.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250226_20250329.L3m.R32.RRS.V3_1.Rrs.4km.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250301_20250331.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250301_20250331.L3m.MO.RRS.V3_1.Rrs.4km.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250305.L3m.DAY.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250305.L3m.DAY.RRS.V3_1.Rrs.4km.nc']]
We can filter/select files with a specific name format using granule_name.
# Level 3 data filtering on file/granule name
results = earthaccess.search_data(
short_name = 'PACE_OCI_L3M_RRS',
temporal = ("2025-03-05", "2025-03-05"),
granule_name="*.MO.*.0p1deg.*"
)
len(results)
1
Refining the search#
Here are the common arguments for earthaccess.search_data():
bounding_boxis a lat/lon box, e.g.(lon_min=-73.5, lat_min=33.5, lon_max=-43.5, lat_max=43.5)temporalis a date range, e.g.("2020-01-16", "2020-12-16")cloud_coveris a range(0, 10)granule_nameis a matching filter, e.g."*.DAY.*.0p1deg.*"short_nameorcollection_id
Summary#
earthaccess.login()to authenticateearthaccess.search_data()to find the data files. This gives an object with all the info needed to get the data.earthaccess.search_data()returns a dictionary/list with information on the data files.
# Level 3 4km global
results = earthaccess.search_data(
short_name = 'PACE_OCI_L3M_RRS',
temporal = ("2025-03-05", "2025-03-05"),
granule_name="*.MO.*.0p1deg.*"
)
# Level 2 1km swath (tiles)
results = earthaccess.search_data(
short_name = 'PACE_OCI_L2_AOP',
temporal = ("2025-03-05", "2025-03-05"),
bounding_box = (-90.0, 40.0, -75.0, 47.0)
)
Load data into memory using xarray#
Following the search for data, we will load data directly into memory using xarray functions.
What are the granules (files)#
# Level 3 data (global)
results = earthaccess.search_data(
short_name = 'PACE_OCI_L3M_RRS',
temporal = ("2025-03-01", "2025-03-01"),
granule_name="*.MO.*.0p1deg.*"
)
len(results)
1
results[0]
Step 1 Set up a file pointer#
We use earthaccess’s open() method to create a ‘fileset’ to the cloud objects with the info needed to open data in cloud buckets. earthaccess does a lot of heavy lifting for us. It identifies the downloadable links, passes our Earthdata Login credentials, and saves the files with the proper names.
fileset = earthaccess.open(results)
Step 2 Read the file using xarray.open_dataset()#
We can load the information about the data and look at its properties without actually loading all the data into memory. Here we load one file using fileset[0].
# Open with xarray
import xarray as xr
ds = xr.open_dataset(fileset[0], chunks={})
ds
<xarray.Dataset> Size: 4GB
Dimensions: (lat: 1800, lon: 3600, wavelength: 172, rgb: 3,
eightbitcolor: 256)
Coordinates:
* lat (lat) float32 7kB 89.95 89.85 89.75 ... -89.75 -89.85 -89.95
* lon (lon) float32 14kB -179.9 -179.9 -179.8 ... 179.8 179.9 180.0
* wavelength (wavelength) float64 1kB 346.0 348.0 351.0 ... 714.0 717.0 719.0
Dimensions without coordinates: rgb, eightbitcolor
Data variables:
Rrs (lat, lon, wavelength) float32 4GB dask.array<chunksize=(16, 1024, 8), meta=np.ndarray>
palette (rgb, eightbitcolor) uint8 768B dask.array<chunksize=(3, 256), meta=np.ndarray>
Attributes: (12/64)
product_name: PACE_OCI.20250301_20250331.L3m.MO.RRS....
instrument: OCI
title: OCI Level-3 Standard Mapped Image
project: Ocean Biology Processing Group (NASA/G...
platform: PACE
source: satellite observations from OCI-PACE
... ...
identifier_product_doi: 10.5067/PACE/OCI/L3M/RRS/3.1
keywords: Earth Science > Oceans > Ocean Optics ...
keywords_vocabulary: NASA Global Change Master Directory (G...
data_bins: 3548336
data_minimum: -0.009993999
data_maximum: 0.09678001We have “lazy” loaded the data, not loaded into memory. This is important since the data are big.
ds_size_gb = ds.nbytes / 1e9
print(f"Dataset size: {ds_size_gb:.2f} GB")
Dataset size: 4.46 GB
Summary#
earthaccess.open()to create an object (fileset or path) for working with data in the cloud.xarray.open_dataset()to open the file (but not load data into memory yet)
# Get results
results = earthaccess.search_data(
short_name = 'PACE_OCI_L3M_RRS',
temporal = ("2025-03-05", "2025-03-05"),
granule_name="*.MO.*.0p1deg.*"
)
# lazy load data
import xarray as xr
fileset = earthaccess.open(results)
ds = xr.open_dataset(fileset[0], chunks={})
ds
In some cases you may want to download your assets. Use the earthaccess.download() function.
downloaded_files = earthaccess.download(
results[0:9],
local_path='data',
)
Example: Level 3 mapped data L3M#
Level 3 data is on a global lat/lon grid. The short name will have L3M in it. The resolution (width/height) of the grids is either 4km or 0.1 degrees.
Let’s load a year of data.
Step 1. Search for the data#
# Level 3 data (global)
results = earthaccess.search_data(
short_name = 'PACE_OCI_L3M_RRS',
temporal = ("2024-03-01", "2025-02-01"),
granule_name="*.MO.*.0p1deg.*"
)
len(results)
12
# look at the file names to make sure they are ok
[res.data_links() for res in results]
[['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20240301_20240331.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20240401_20240430.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20240501_20240531.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20240601_20240630.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20240701_20240731.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20240801_20240831.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20240901_20240930.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20241001_20241031.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20241101_20241130.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20241201_20241231.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250101_20250131.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc'],
['https://obdaac-tea.earthdatacloud.nasa.gov/ob-cumulus-prod-public/PACE_OCI.20250201_20250228.L3m.MO.RRS.V3_1.Rrs.0p1deg.nc']]
Step 2. Set up the fileset#
This step is needed so that xarray knows how to interact with the data in the cloud bucket.
fileset = earthaccess.open(results)
Step 3. Read one file with xarray#
Open one file using open_dataset(). Notice the coordinates (lat/lon and wavelength) and the data variables (Rrs).
ds1 = xr.open_dataset(fileset[1])
ds1
<xarray.Dataset> Size: 4GB
Dimensions: (lat: 1800, lon: 3600, wavelength: 172, rgb: 3,
eightbitcolor: 256)
Coordinates:
* lat (lat) float32 7kB 89.95 89.85 89.75 ... -89.75 -89.85 -89.95
* lon (lon) float32 14kB -179.9 -179.9 -179.8 ... 179.8 179.9 180.0
* wavelength (wavelength) float64 1kB 346.0 348.0 351.0 ... 714.0 717.0 719.0
Dimensions without coordinates: rgb, eightbitcolor
Data variables:
Rrs (lat, lon, wavelength) float32 4GB ...
palette (rgb, eightbitcolor) uint8 768B ...
Attributes: (12/64)
product_name: PACE_OCI.20240401_20240430.L3m.MO.RRS....
instrument: OCI
title: OCI Level-3 Standard Mapped Image
project: Ocean Biology Processing Group (NASA/G...
platform: PACE
source: satellite observations from OCI-PACE
... ...
identifier_product_doi: 10.5067/PACE/OCI/L3M/RRS/3.1
keywords: Earth Science > Oceans > Ocean Optics ...
keywords_vocabulary: NASA Global Change Master Directory (G...
data_bins: 3355114
data_minimum: -0.009991998
data_maximum: 0.099733464# here are the wavelengths
ds1['wavelength']
<xarray.DataArray 'wavelength' (wavelength: 172)> Size: 1kB
array([346., 348., 351., 353., 356., 358., 361., 363., 366., 368., 371., 373.,
375., 378., 380., 383., 385., 388., 390., 393., 395., 398., 400., 403.,
405., 408., 410., 413., 415., 418., 420., 422., 425., 427., 430., 432.,
435., 437., 440., 442., 445., 447., 450., 452., 455., 457., 460., 462.,
465., 467., 470., 472., 475., 477., 480., 482., 485., 487., 490., 492.,
495., 497., 500., 502., 505., 507., 510., 512., 515., 517., 520., 522.,
525., 527., 530., 532., 535., 537., 540., 542., 545., 547., 550., 553.,
555., 558., 560., 563., 565., 568., 570., 573., 575., 578., 580., 583.,
586., 588., 613., 615., 618., 620., 623., 625., 627., 630., 632., 635.,
637., 640., 641., 642., 643., 645., 646., 647., 648., 650., 651., 652.,
653., 655., 656., 657., 658., 660., 661., 662., 663., 665., 666., 667.,
668., 670., 671., 672., 673., 675., 676., 677., 678., 679., 681., 682.,
683., 684., 686., 687., 688., 689., 691., 692., 693., 694., 696., 697.,
698., 699., 701., 702., 703., 704., 706., 707., 708., 709., 711., 712.,
713., 714., 717., 719.])
Coordinates:
* wavelength (wavelength) float64 1kB 346.0 348.0 351.0 ... 714.0 717.0 719.0
Attributes:
long_name: wavelengths
units: nm
valid_min: 0
valid_max: 20000Plot#
We can plot this object. But let’s not plot the full global data as that is 4+ Gb. We will plot a subset using the slice() function. This way we only load the data needed to plot into memory. We need to
pick the variable. Let’s use
Rrspick a wavelength using
sel()slice to a smaller lat/lon using
slice()plot
# Plot
rrs = ds1['Rrs']
rrs346 = rrs.sel(wavelength=346)
rrs346_sub = rrs346.sel(lat=slice(47, 40), lon=slice(-90, -75))
rrs346_sub.plot(robust=True);
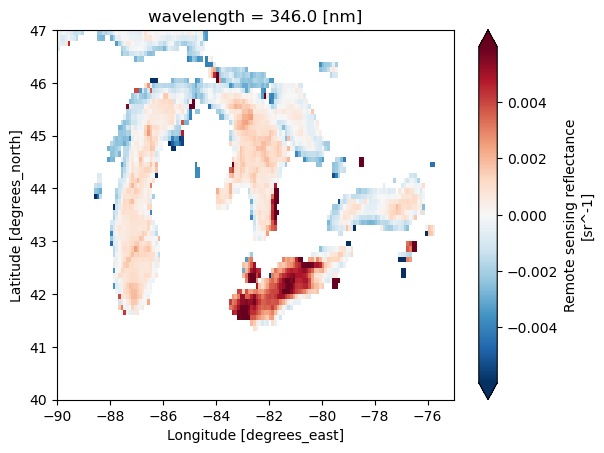
# Note this is how we'd usually write this
rrs = ds["Rrs"].sel(
wavelength=346,
lat=slice(47, 40),
lon=slice(-90, -75)
)
rrs.plot(robust=True);
Summary#
For Level 3, we use the mapped data product, L3M and the steps to open with xarray are
earthaccess.search_data()withgranule_nameto filter to the temporal binning (day, 8-day, month) and resolution (4km or 0.1 degree). We do not needbounding_boxsince level 3 is global.earthaccess.open()to open the search resultsds=xr.open_dataset(fileset[i])to open the i-th file in the fileset ords=xr.open_mfdataset(fileset)to open multiple datasets.
results = earthaccess.search_data(
short_name = 'PACE_OCI_L3M_RRS',
temporal = ("2024-03-01", "2025-02-01"),
granule_name="*.MO.*.0p1deg.*"
)
fileset = earthaccess.open(results)
ds = xr.open_dataset(fileset[1])
Optional. Open all 12 months and make a plot#
Open all 12 files with open_mfdataset() and since the netcdf files do not have a time coordinate (just lat, lon, wavelength), we need to tell xarray to combine on a new dimension “time”.
# We can open all the files but note there is no time coordinate so we need
# combine="nested" and concat_dim
ds = xr.open_mfdataset(
fileset,
combine="nested",
concat_dim="time")
ds
<xarray.Dataset> Size: 53GB
Dimensions: (time: 12, lat: 1800, lon: 3600, wavelength: 172, rgb: 3,
eightbitcolor: 256)
Coordinates:
* wavelength (wavelength) float64 1kB 346.0 348.0 351.0 ... 714.0 717.0 719.0
* lat (lat) float32 7kB 89.95 89.85 89.75 ... -89.75 -89.85 -89.95
* lon (lon) float32 14kB -179.9 -179.9 -179.8 ... 179.8 179.9 180.0
Dimensions without coordinates: time, rgb, eightbitcolor
Data variables:
Rrs (time, lat, lon, wavelength) float32 53GB dask.array<chunksize=(1, 16, 1024, 8), meta=np.ndarray>
palette (time, rgb, eightbitcolor) uint8 9kB dask.array<chunksize=(1, 3, 256), meta=np.ndarray>
Attributes: (12/64)
product_name: PACE_OCI.20240301_20240331.L3m.MO.RRS....
instrument: OCI
title: OCI Level-3 Standard Mapped Image
project: Ocean Biology Processing Group (NASA/G...
platform: PACE
source: satellite observations from OCI-PACE
... ...
identifier_product_doi: 10.5067/PACE/OCI/L3M/RRS/3.1
keywords: Earth Science > Oceans > Ocean Optics ...
keywords_vocabulary: NASA Global Change Master Directory (G...
data_bins: 3315440
data_minimum: -0.009997999
data_maximum: 0.09494999# Let's add the time coord since we will likely want to subset on time later
import pandas as pd
t = pd.date_range(start="2024-03-01", end="2025-02-01", freq="MS")
ds = ds.assign_coords(time=t)
ds
<xarray.Dataset> Size: 53GB
Dimensions: (time: 12, lat: 1800, lon: 3600, wavelength: 172, rgb: 3,
eightbitcolor: 256)
Coordinates:
* wavelength (wavelength) float64 1kB 346.0 348.0 351.0 ... 714.0 717.0 719.0
* lat (lat) float32 7kB 89.95 89.85 89.75 ... -89.75 -89.85 -89.95
* lon (lon) float32 14kB -179.9 -179.9 -179.8 ... 179.8 179.9 180.0
* time (time) datetime64[ns] 96B 2024-03-01 2024-04-01 ... 2025-02-01
Dimensions without coordinates: rgb, eightbitcolor
Data variables:
Rrs (time, lat, lon, wavelength) float32 53GB dask.array<chunksize=(1, 16, 1024, 8), meta=np.ndarray>
palette (time, rgb, eightbitcolor) uint8 9kB dask.array<chunksize=(1, 3, 256), meta=np.ndarray>
Attributes: (12/64)
product_name: PACE_OCI.20240301_20240331.L3m.MO.RRS....
instrument: OCI
title: OCI Level-3 Standard Mapped Image
project: Ocean Biology Processing Group (NASA/G...
platform: PACE
source: satellite observations from OCI-PACE
... ...
identifier_product_doi: 10.5067/PACE/OCI/L3M/RRS/3.1
keywords: Earth Science > Oceans > Ocean Optics ...
keywords_vocabulary: NASA Global Change Master Directory (G...
data_bins: 3315440
data_minimum: -0.009997999
data_maximum: 0.09494999Let’s plot all the months and fix the warping of space by adding a lat/lon projection, aka tell the plotting that this is lat/lon on a globe.
rrs = ds["Rrs"].sel(
wavelength=346,
lat=slice(47, 40),
lon=slice(-90, -75)
)
import cartopy.crs as ccrs
rrs.plot(
col="time", # one panel per month
col_wrap=4, # 4 columns per row
robust=True, # ignore outliers for color scale
figsize=(12, 4),
subplot_kws={"projection": ccrs.PlateCarree()},
transform=ccrs.PlateCarree()
)
<xarray.plot.facetgrid.FacetGrid at 0x7f413816ec90>
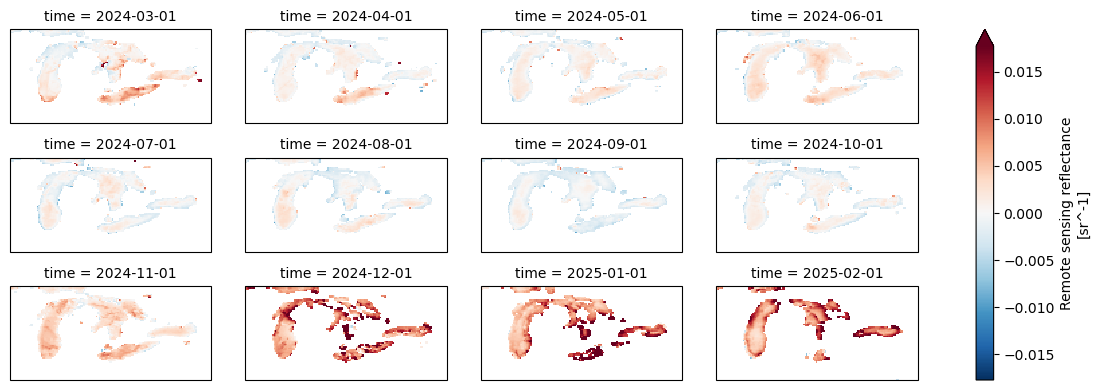
Level 2 data#
Level 2 data is not on a global lat/lon grid. Instead each swath (satelite path) has “lines” along the path and “pixels” across the lines (perpendicular to path). In addition, the data files are GROUPED netcdfs (different format). We need to open these with open_datatree and then merge all the groups.
Let’s get the swath(s) over the U.S. Great Lakes on March 5th, 2025.
import xarray as xr
import earthaccess
auth = earthaccess.login(persist=True)
results = earthaccess.search_data(
short_name = 'PACE_OCI_L2_AOP',
temporal = ("2025-03-05", "2025-03-05"),
bounding_box = (-90.0, 40.0, -75.0, 47.0)
)
len(results) # one of this is a monthly average
3
# Step 1 set up the fileset
fileset = earthaccess.open(results)
# Step 2 open L2 swath files with xarray open_datatree
datatree = xr.open_datatree(fileset[1], decode_timedelta=False)
datatree.groups
('/',
'/sensor_band_parameters',
'/scan_line_attributes',
'/geophysical_data',
'/navigation_data',
'/processing_control',
'/processing_control/input_parameters',
'/processing_control/flag_percentages')
The geophysical data is the data like Rrs and the navigation data has the lat/lon information for each line/pixel.
# Step 3 merge the groups all together.
# This works due to the values in each group being the same shape.
ds_l2 = xr.merge(datatree.to_dict().values())
ds_l2 = ds_l2.set_coords(("longitude", "latitude"))
ds_l2
<xarray.Dataset> Size: 3GB
Dimensions: (number_of_bands: 286, number_of_reflective_bands: 286,
wavelength_3d: 172, number_of_lines: 1710,
pixels_per_line: 1272)
Coordinates:
* wavelength_3d (wavelength_3d) float64 1kB 346.0 348.0 351.0 ... 717.0 719.0
longitude (number_of_lines, pixels_per_line) float32 9MB ...
latitude (number_of_lines, pixels_per_line) float32 9MB ...
Dimensions without coordinates: number_of_bands, number_of_reflective_bands,
number_of_lines, pixels_per_line
Data variables: (12/30)
wavelength (number_of_bands) float64 2kB ...
vcal_gain (number_of_reflective_bands) float32 1kB ...
vcal_offset (number_of_reflective_bands) float32 1kB ...
F0 (number_of_reflective_bands) float32 1kB ...
aw (number_of_reflective_bands) float32 1kB ...
bbw (number_of_reflective_bands) float32 1kB ...
... ...
aot_865 (number_of_lines, pixels_per_line) float32 9MB ...
angstrom (number_of_lines, pixels_per_line) float32 9MB ...
avw (number_of_lines, pixels_per_line) float32 9MB ...
nflh (number_of_lines, pixels_per_line) float32 9MB ...
l2_flags (number_of_lines, pixels_per_line) int32 9MB ...
tilt (number_of_lines) float32 7kB ...
Attributes: (12/47)
title: OCI Level-2 Data AOP
product_name: PACE_OCI.20250305T174755.L2.OC_AOP.V3_...
processing_version: 3.1
history: l2gen par=/data3/sdpsoper/vdc/vpu22/wo...
instrument: OCI
platform: PACE
... ...
geospatial_lon_min: -102.731316
startDirection: Ascending
endDirection: Ascending
day_night_flag: Day
earth_sun_distance_correction: 1.0163003206253052
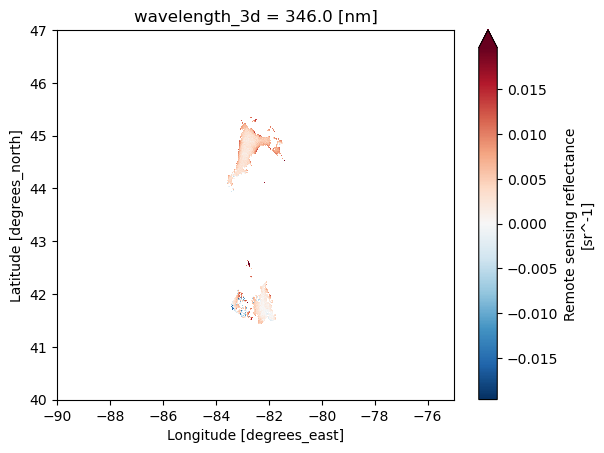
geospatial_bounds: POLYGON ((-63.58643 55.76101, -102.731...Let’s plot. Hmm it’s mainly blank. This is daily swath data. There were lots of clouds that day or the swath only included part of region.
Specify the variable
Rrsin[ ]Spefify the wavelength, a coordinate, using
.sel()Plot and tell it to use longitude for x and latitude for y.
# Plot
rrs = ds_l2['Rrs'].sel(
wavelength_3d=346)
rrs.plot(robust=True,
x="longitude", y="latitude",
xlim=(-90, -75), ylim=(40, 47));
Optional. Plot with the outline of the Great Lakes#
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
import cartopy.feature as cfeature
fig = plt.figure(figsize=(6,5))
ax = plt.axes(projection=ccrs.PlateCarree())
ax.set_extent([-90, -75, 40, 47], crs=ccrs.PlateCarree())
ax.add_feature(cfeature.LAKES.with_scale("50m"),
facecolor="none", edgecolor="grey", linewidth=0.8)
# swath pieces
da = ds_l2["Rrs"].sel(wavelength_3d=346) # (lines, pixels)
lat = ds_l2["latitude"]
lon = ds_l2["longitude"]
# add to your existing ax (before plt.show)
im = ax.pcolormesh(
lon, lat, da,
transform=ccrs.PlateCarree(),
shading="nearest", # lighter than 'auto'
rasterized=True
)
plt.colorbar(im, ax=ax, label="Rrs")
plt.show()
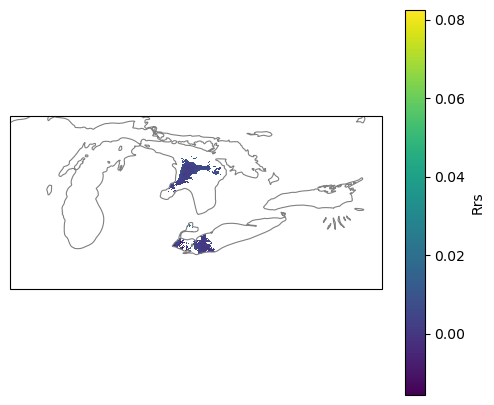
Summary#
With level 2 data, we need to use xarray.open_datatree() and then do a merge step to get all the variables in the groups. Note we can only work with one level 2 dataset at a time.
res=earthaccess.search_data()withbounding_boxto filter to the region of interest.fileset=earthaccess.open(res)datatree=xarray.open_datatree(fileset[i])to open the i-th file in the fileset.ds=xarray.merge(datatree.to_dict().values())to merge all the groupsds = ds.set_coords(("longitude", "latitude"))to make longitude and latitude coords (will help with plotting.
# Get L2 data
results = earthaccess.search_data(
short_name = 'PACE_OCI_L2_AOP',
temporal = ("2025-03-05", "2025-03-05"),
bounding_box = (-90.0, 40.0, -75.0, 47.0)
)
fileset = earthaccess.open(results)
# Open with the grouped netCDFs with open_datatree
datatree = xr.open_datatree(fileset[1], decode_timedelta=False)
ds_l2 = xr.merge(datatree.to_dict().values())
ds_l2 = ds_l2.set_coords(("longitude", "latitude"))
ds_l2
Working with larger than memory data#
Here are a few basics to know when working with larger-than memory data.
Chunks#
The PACE data are cloud-optimized (read optimized) so that we don’t have to read the whole file into memory to work with it and we don’t have to download the whole thing if we don’t need to.
import earthaccess
import xarray as xr
results = earthaccess.search_data(
short_name = 'PACE_OCI_L3M_RRS',
temporal = ("2024-03-01", "2025-04-01"),
granule_name="*.MO.*.0p1deg.*"
)
fileset = earthaccess.open(results)
# this creates a chunked reference to our data that we can load chunk by chunk
ds_chunk = xr.open_dataset(fileset[0], chunks={})
# this creates a reference that we have to load in its entirety
ds = xr.open_dataset(fileset[0])
ds_size_gb = ds.nbytes / 1e9
print(f"Dataset size: {ds_size_gb:.2f} GB")
Dataset size: 4.46 GB
# Check that ds_chunk is chunked
ds_chunk.chunks
Frozen({'lat': (16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 8), 'lon': (1024, 1024, 1024, 528), 'wavelength': (8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 8, 4), 'rgb': (3,), 'eightbitcolor': (256,)})
# but ds is not chunked
ds.chunks
Frozen({})
Chunking allows operations on bigger than memory datasets#
Chunking let’s us work with big datasets without blowing up our memory. This task takes awhile but it doesn’t blow up our little 2Gb RAM virtual machines. xarray and dask are smart and work through the task chunk by chunk. Downside is that the chunked dataset is chunked into little chunks which mean a lot of queries to the cloud data to read each little chunk. That means it will be very slow.
%%time
# ds is not chunked
# This loads a 200 Mb array at once. Watch below how the memory increases (if on a JupyterHub)
var = ds["Rrs"].isel(wavelength=slice(1,9))
global_mean = var.mean().values
CPU times: user 15.3 s, sys: 7.04 s, total: 22.4 s
Wall time: 1min 13s
%%time
# ds_chunk is chunked into little chunks
# This can be done chunk by chunk and will increase the memory usage much since chunks are tiny, but slow.
var = ds_chunk["Rrs"].isel(wavelength=slice(1,9))
global_mean = var.mean().values
CPU times: user 7.2 s, sys: 1.19 s, total: 8.39 s
Wall time: 2min 21s
Working with Larger-Than-Memory Data When You Cannot Change Chunk Sizes#
In the case of the PACE data, the chunk sizes are small and computations are much slower than if we could load all the data into memory but we don’t blow up the memory so we can work with big files on machines with minimal RAM or with lots of little dask workers in a big RAM machine.
When datasets are stored in the cloud with very small physical chunks, the key is to design computations that minimize repeated remote reads and avoid loading the full array into RAM. Even though the tiny chunking slows down global operations, Dask + Xarray can still handle larger-than-memory workflows efficiently if you structure your analysis around streaming, subsetting, and locality or we can use a non-chunked dataset if we only load in smaller than memory subsets.
Practical Tips#
Always subset to your region and time of interest before working with your dataset. So
sel()andisel()beforemean()orvar(). Focus only on the wavelengths and variables of interest (so use likeds["Rrs"]to get just that variable).Avoid operations that require full materialization such as
.load()or.valueson large arrays. Instead, keep arrays lazy and call.compute()only on reduced results (e.g., monthly means, statistics, anomalies), which keeps memory usage bounded.Use
xarray.open_data(..., chunks={})to create a version of your data that has the chunk references if you need to make sure memory use is limited. This is going to be slower, but won’t blow up memory. Useds.chunks()on your xarray dataset to find out if the dataset is chunked or not..plot()will load all the data so make sure to plot smaller regions.hvplot()uses dask if you want to plot something that is larger than memory.Coding LLMs (Claude, ChatGPT) are very helpful in debugging memory issues and chunking issues.
If you need to run computations with dask, use a machine with big memory and lots of workers because your chunks are small and each worker doesn’t need too much memory. But you will likely be limited by the number of http connections you can make (since the bottleneck in cloud data is often the I/O calls).
Summary#
This concludes the earthaccess tutorial for PACE data.
Opening Level 3 netcdfs with xarray
Opening Level 2 netcdfs with xarray and datatree
Working with larger than memory data.




