OOI Arrays: Overview and Context for CHLA Profiling#
Author: Eli Holmes (NOAA) Last updated: Jan 29, 2026
The Ocean Observatories Initiative (OOI) is a long-term, NSF-funded ocean observing program designed to provide continuous, high-resolution measurements across key regions of the global and coastal ocean.
OOI systems collect physical, chemical, biological, and geological observations using a combination of:
Moorings: fixed location (surface, mid-water, deep-water) — These are sensors at fixed discrete depths. CHLA is available only at those specific depths.
Depth Profiler systems: fixed location
Wire-following profilers (WFPs) — Provide high-resolution vertical depth profiles, sampling repeatedly up and down the water column.
Shallow profilers (SPs) — Motorized profilers delivering dense CHLA depth profiles in the 0-200m range, often with higher sampling frequency than WFPs.
Deep profilers (DPs) — Provide deep water (> 200m) vertical profiles.
Autonomous gliders: moving location — Mobile platforms that execute repeated sawtooth depth profiles and produce full vertical CHLA measurements for each dive.
Across these platforms, OOI instruments collect high-frequency, vertically resolved biological measurements, including: chlorophyll-a concentration, particulate organic matter & phytoplankton biomass, colored dissolved organic matter (CDOM), photosynthetically available radiation (PAR), sea water physical properties (temperature, salinity, and density), dissolved oxygen, nitrate, pH, pCO₂. We are interested in modeling CHLA vertical depth profiles and will focus on the chlorophyll-a concentration measurement. OOI data is openly available through the OOI Data Portal and ERDDAP servers.
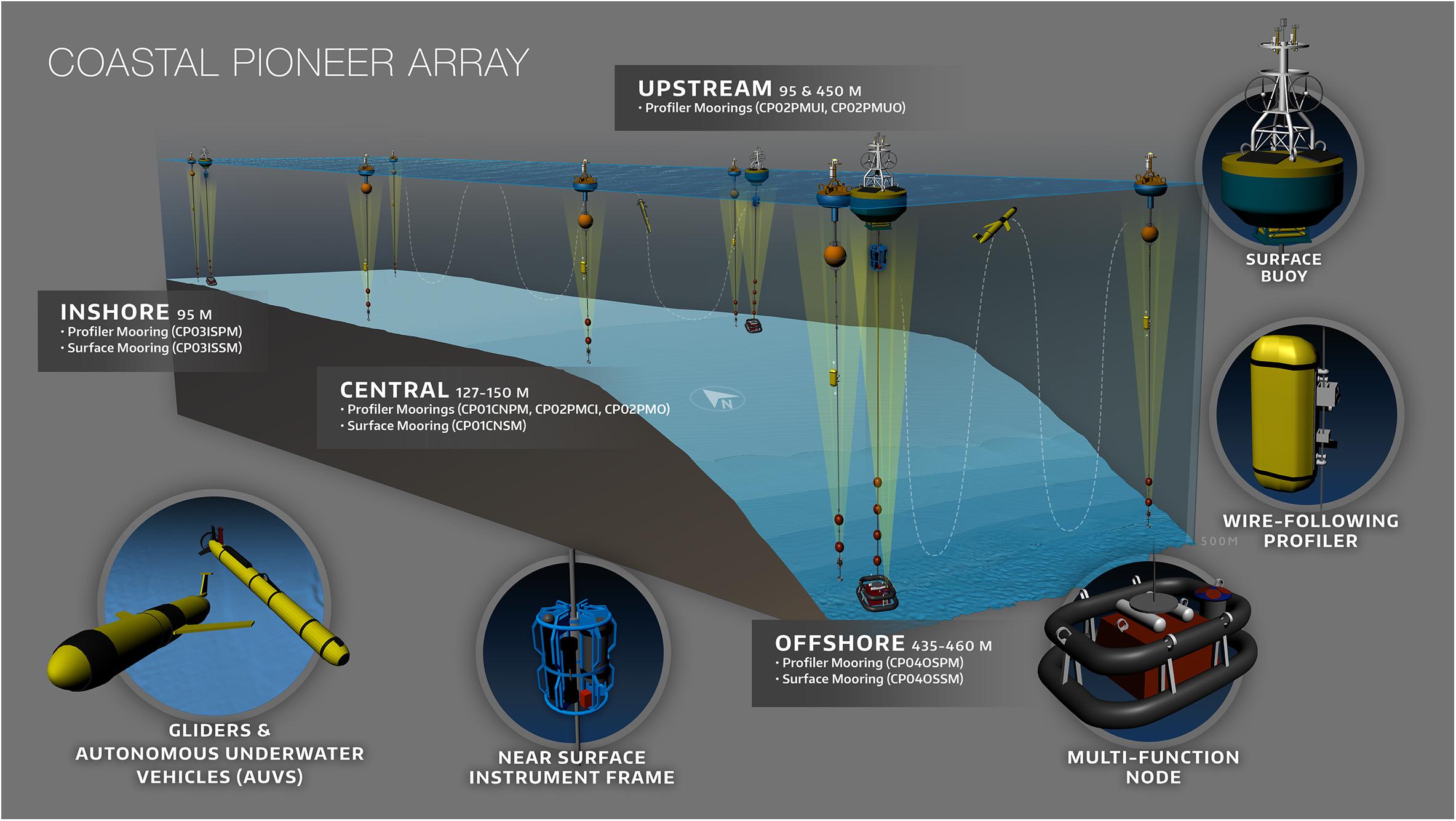
OOI Regional Arrays#
OOI consists of two major coastal arrays, a regional cabled array, and four global open-ocean arrays. Coastal arrays sample continental shelf, slope, and upwelling environments that exhibit strong seasonal biological and physical variability. These sites typically include vertically profiling platforms that provide full or partial CHLA depth profiles in the upper 0–200 m. Global Arrays observe key climate-relevant open-ocean regions and generally do not include vertical profilers, but they do provide CHLA at fixed depths within the 0–200 m zone, which can still be mapped into depth bins.
Coastal Stations:
Endurance Array (Oregon & Washington) — Samples an eastern boundary upwelling system across nearshore, shelf, and offshore environments. Platforms include surface and subsurface moorings (fixed-depth CHLA), wire-following profilers (0–200 m), cabled shallow/deep profilers, and autonomous gliders (0–200+ m profiles).
Pioneer Array (Northwest Atlantic / Mid-Atlantic Bight) — Observes cross-shelf and shelf-break frontal systems with strong biological gradients. Platforms include multi-tier moorings, surface buoys, gliders, and (in most years) shallow profilers that provided high-resolution 0–200 m CHLA profiles.
Regional Cabled Array (Washington–Oregon offshore) — Provides real-time power and data to offshore slope and deepwater nodes. Only the cabled shallow profilers contribute 0–200 m CHLA profiles; most deep or seafloor instruments do not sample CHLA.
Global Stations:
Papa Array (Gulf of Alaska) — Subarctic gyre; CHLA sensors at fixed depths (typically 7–50 m).
Irminger Sea Array (Greenland–Iceland region) — High-latitude convection region; CHLA at fixed depths in the upper 100 m.
Argentine Basin Array (Southwest Atlantic) — Deep-ocean circulation site; limited CHLA at one or two fixed depths (~30 m).
Southern Ocean Array (South of Tasmania) — High-mixing Southern Ocean environment; CHLA measurements at fixed depths in the upper 50 m.
Workflow#
This notebook processes CHLA data from OOI platforms showing:
automated ERDDAP retrieval,
daily depth-binned CHLA profiles,
merged datasets across platforms and regions
at the end, I show how I did the matchups
We work with a set of Endurance platforms listed in endurance_dict, including:
Offshore and shelf surface moorings (near-surface instrument frames with fluorometers)
Cabled shallow and deep profilers
Wire-following profilers on offshore moorings
Inshore and shelf surface-piercing profilers
For each platform, we:
Request CHLA data from the OOI ERDDAP server.
Process the data into daily mean CHLA profiles in fixed 10 m depth bins between 0 and 200 m.
Save per-platform parquet files and then merge them into a single, combined Endurance dataset.
Plot example CHLA–depth profiles for individual daily “profiles” (platform × day).
Inspect ERDDAP variables for a single instrument#
First, inspect the available variables for one representative dataset on the OOI ERDDAP server.
This helps confirm the exact variable names for chlorophyll-a and related quality-control flags before building generalized code.
# See the variables
import pandas as pd
base = "https://erddap.dataexplorer.oceanobservatories.org/erddap/tabledap"
dataset_id = "ooi-ce04osps-sf01b-3a-flortd104"
header_url = f"{base}/{dataset_id}.csv?"
# read only the header, skip the units row
df0 = pd.read_csv(header_url, skiprows=[1], nrows=0)
print(df0.columns)
Prototype daily CHLA binning and profile plotting for one mooring#
Here we prototype the workflow for one Endurance instrument:
Request time series from ERDDAP.
Convert depth from
z(altitude, negative down) to positive meters.Filter to the 0–200 m range.
Bin CHLA into 10 m depth bins (0–10, 10–20, …, 190–200 m).
Aggregate to daily means per depth bin.
Pick one daily profile and plot CHLA vs depth.
# See a dataframe with the variables we want
import pandas as pd
import os
import numpy as np
dataset_id = "ooi-ce04osps-sf01b-3a-flortd104"
chl_var = "mass_concentration_of_chlorophyll_a_in_sea_water_profiler_depth_enabled"
url = (
"https://erddap.dataexplorer.oceanobservatories.org/erddap/tabledap/"
f"{dataset_id}.csv?"
"time%2Clatitude%2Clongitude%2Cz%2C"
f"{chl_var}%2C"
f"{chl_var}_qc_agg"
"&time%3E=2024-03-01"
"&time%3C=2025-12-05T00%3A00%3A00Z"
"&z%3E=-200"
)
df = pd.read_csv(url, skiprows=[1])
df["depth"] = -df["z"]
qc_col = "mass_concentration_of_chlorophyll_a_in_sea_water_profiler_depth_enabled_qc_agg"
mask = (df["depth"] >= 0) & (df["depth"] < 200) & (df[qc_col] == 1.0)
df_good = df[mask].copy()
df_good
| time | latitude | longitude | z | mass_concentration_of_chlorophyll_a_in_sea_water_profiler_depth_enabled | mass_concentration_of_chlorophyll_a_in_sea_water_profiler_depth_enabled_qc_agg | depth | |
|---|---|---|---|---|---|---|---|
| 0 | 2024-03-01T00:00:00Z | 44.369353 | -124.954108 | -161.0 | 0.033150 | 1 | 161.0 |
| 1 | 2024-03-01T00:01:00Z | 44.369353 | -124.954108 | -161.0 | 0.030857 | 1 | 161.0 |
| 2 | 2024-03-01T00:02:00Z | 44.369353 | -124.954108 | -161.0 | 0.037528 | 1 | 161.0 |
| 3 | 2024-03-01T00:03:00Z | 44.369353 | -124.954108 | -161.0 | 0.038779 | 1 | 161.0 |
| 4 | 2024-03-01T00:04:00Z | 44.369353 | -124.954108 | -161.0 | 0.022100 | 1 | 161.0 |
| ... | ... | ... | ... | ... | ... | ... | ... |
| 2350658 | 2025-12-04T23:56:00Z | 44.369353 | -124.954108 | -192.0 | 0.047337 | 1 | 192.0 |
| 2350659 | 2025-12-04T23:57:00Z | 44.369353 | -124.954108 | -192.0 | 0.046868 | 1 | 192.0 |
| 2350660 | 2025-12-04T23:58:00Z | 44.369353 | -124.954108 | -192.0 | 0.049680 | 1 | 192.0 |
| 2350661 | 2025-12-04T23:59:00Z | 44.369353 | -124.954108 | -192.0 | 0.049446 | 1 | 192.0 |
| 2350662 | 2025-12-05T00:00:00Z | 44.369353 | -124.954108 | -192.0 | 0.054280 | 1 | 192.0 |
2034006 rows × 7 columns
Define a reusable function to fetch and bin CHLA from ERDDAP#
Next we wrap the ERDDAP access and daily-binning logic into a reusable helper:
ooi_mooring_save_file(url, description, id, save, out_dir)
This function:
Downloads CHLA (and QC flags, when available) from ERDDAP for a given dataset.
Converts depths to positive meters and filters to 0–200 m.
Computes daily mean CHLA in 10 m depth bins.
Ensures all CHLA depth-bin columns (e.g.,
CHLA_0_10, …,CHLA_190_200) are present.Adds metadata such as latitude, longitude, description, and dataset ID.
Optionally saves the daily profiles to a parquet file for that platform.
# Get and save a file
import numpy as np
import pandas as pd
from pathlib import Path
from urllib.parse import urlparse
import os
def ooi_mooring_save_file(url, description=None, chl_var="mass_concentration_of_chlorophyll_a_in_sea_water_profiler_depth_enabled",
id=None, save=False, out_dir=Path("_temp_data/ooi")):
df = pd.read_csv(url, skiprows=[1])
qc_col = f"{chl_var}_qc_agg"
# Rename CHLA columns
df = df.rename(columns={
chl_var: "CHLA",
})
chla_col = "CHLA"
# Time + date
df["time"] = pd.to_datetime(df["time"])
df["date"] = df["time"].dt.floor("D")
# Depth (positive down)
df["depth"] = -df["z"]
# Filter 0–200 m and QC
mask = (df["depth"] >= 0) & (df["depth"] < 200) & (df[qc_col] == 1.0)
df_good = df[mask].copy()
# 10 m bins
bins = np.arange(0, 210, 10) # 0,10,...,200
labels = [f"{b}-{b+10}" for b in bins[:-1]]
df_good["depth_bin"] = pd.cut(
df_good["depth"],
bins=bins,
labels=labels,
right=False,
)
# ---- PER-DAY × DEPTH-BIN AGGREGATION (with lat/lon) ----
grp = (
df_good
.groupby(["date", "depth_bin"], observed=True)
.agg(
lat=("latitude", "first"),
lon=("longitude", "first"),
chla=(chla_col, "mean"),
)
.reset_index()
)
# ---- PIVOT TO WIDE FORM ----
wide = grp.pivot(
index=["date", "lat", "lon"],
columns="depth_bin",
values="chla",
).reset_index()
# Rename depth-bin columns "0-10" → "chla_0_10"
wide.columns = [
("CHLA_" + str(c).replace("-", "_")) if c not in ("date", "lat", "lon") else c
for c in wide.columns
]
# rename date → time
wide = wide.rename(columns={"date": "time"})
if description is not None:
wide["description"]=description
if save:
parsed = urlparse(url)
path = parsed.path
filename = os.path.basename(path) # 'ooi-ce04osps-sf01b-3a-flortd104.csv'
dataset_id = filename.split('.')[0] # 'ooi-ce04osps-sf01b-3a-flortd104'
out_dir.mkdir(parents=True, exist_ok=True)
out_path = f"{out_dir}/{dataset_id}.parquet"
wide.to_parquet(out_path)
return wide, out_path
return wide, None
Define OOI platforms of interest#
Here we list the OOI platforms we want to process.
Each key in ooi_dict is an ERDDAP dataset_id on https://erddap.dataexplorer.oceanobservatories.org/erddap, and each value is a human-readable description of the mooring or profiler (location and instrument type, e.g., 3-wavelength fluorometer).
# The files we want
# to do. add doi's from this: https://oceanobservatories.org/site-list/
ooi_dict = {
"ooi-ce04ossm-rid27-02-flortd000": "Coastal Endurance: Oregon Offshore Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-ce04osps-sf01b-3a-flortd104": "Coastal Endurance: Oregon Offshore Cabled Shallow Profiler Mooring: Shallow Profiler (SF01B): 3-Wavelength Fluorometer",
"ooi-ce09ospm-wfp01-04-flortk000": "Coastal Endurance: Washington Offshore Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-ce01issp-sp001-08-flortj000": "Coastal Endurance: Oregon Inshore Surface Piercing Profiler Mooring: Surface Piercing Profiler: 3-Wavelength Fluorometer",
"ooi-ce04ospd-dp01b-04-flntua103": "Coastal Endurance: Oregon Offshore Cabled Deep Profiler Mooring: Wire-Following Profiler (DP01B): 2-Wavelength Fluorometer",
"ooi-ce02shsp-sp001-07-flortj000": "Coastal Endurance: Oregon Shelf Surface Piercing Profiler Mooring: Surface Piercing Profiler: 3-Wavelength Fluorometer",
"ooi-ce02shsp-sp002-07-flortj000": "Coastal Endurance: Oregon Shelf Surface Piercing Profiler Mooring: Surface Piercing Profiler: 3-Wavelength Fluorometer B",
"ooi-ce06issp-sp001-08-flortj000": "Coastal Endurance: Washington Inshore Surface Piercing Profiler Mooring: Surface Piercing Profiler: 3-Wavelength Fluorometer",
"ooi-ce07shsp-sp001-07-flortj000": "Coastal Endurance: Washington Shelf Surface Piercing Profiler Mooring: Surface Piercing Profiler: 3-Wavelength Fluorometer",
"ooi-cp10cnsm-rid27-02-flortd000": "Coastal Pioneer MAB: Central Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-cp13eapm-wfp01-04-flortk000": "Coastal Pioneer MAB: Eastern Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp14nepm-wfp01-04-flortk000": "Coastal Pioneer MAB: Northeastern Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp13nopm-wfp01-04-flortk000": "Coastal Pioneer MAB: Northern Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp11nosm-rid27-02-flortd000": "Coastal Pioneer MAB: Northern Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-cp14sepm-wfp01-04-flortk000": "Coastal Pioneer MAB: Southeastern Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp13sopm-wfp01-04-flortk000": "Coastal Pioneer MAB: Southern Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp11sosm-rid27-02-flortd000": "Coastal Pioneer MAB: Southern Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-cp02pmci-wfp01-04-flortk000": "Coastal Pioneer NES: Central Inshore Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp02pmco-wfp01-04-flortk000": "Coastal Pioneer NES: Central Offshore Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp01cnpm-wfp01-04-flortk000": "Coastal Pioneer NES: Central Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp01cnsm-rid27-02-flortd000": "Coastal Pioneer NES: Central Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-cp01cnsp-sp001-09-flortj000": "Coastal Pioneer NES: Central Surface Piercing Profiler Mooring: Surface Piercing Profiler: 3-Wavelength Fluorometer",
"ooi-cp03ispm-wfp01-04-flortk000": "Coastal Pioneer NES: Inshore Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp03issm-rid27-02-flortd000": "Coastal Pioneer NES: Inshore Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-cp03issp-sp001-09-flortj000": "Coastal Pioneer NES: Inshore Surface Piercing Profiler Mooring: Surface Piercing Profiler: 3-Wavelength Fluorometer",
"ooi-cp04ospm-wfp01-04-flortk000": "Coastal Pioneer NES: Offshore Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp04ossm-rid27-02-flortd000": "Coastal Pioneer NES: Offshore Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-cp02pmui-wfp01-04-flortk000": "Coastal Pioneer NES: Upstream Inshore Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-cp02pmuo-wfp01-04-flortk000": "Coastal Pioneer NES: Upstream Offshore Profiler Mooring: Wire-Following Profiler: 3-Wavelength Fluorometer",
"ooi-rs03axps-pc03a-4c-flordd303": "Regional Cabled Array: Axial Base Shallow Profiler Mooring: 200m Platform (PC03A): 2-Wavelength Fluorometer",
"ooi-rs03axps-sf03a-3a-flortd301": "Regional Cabled Array: Axial Base Shallow Profiler Mooring: Shallow Profiler (SF03A): 3-Wavelength Fluorometer",
"ooi-rs01sbpd-dp01a-04-flntua102": "Regional Cabled Array: Oregon Slope Base Deep Profiler Mooring: Wire-Following Profiler (DP01A): 2-Wavelength Fluorometer",
"ooi-rs01sbps-pc01a-4c-flordd103": "Regional Cabled Array: Oregon Slope Base Shallow Profiler Mooring: 200m Platform (PC01A): 2-Wavelength Fluorometer",
"ooi-rs01sbps-sf01a-3a-flortd101": "Regional Cabled Array: Oregon Slope Base Shallow Profiler Mooring: Shallow Profiler (SF01A): 3-Wavelength Fluorometer",
"ooi-gp02hypm-wfp03-01-flordl000": "Global Station Papa: Apex Profiler Mooring: Wire-Following Profiler Lower: 2-Wavelength Fluorometer",
"ooi-gp02hypm-wfp02-01-flordl000": "Global Station Papa: Apex Profiler Mooring: Wire-Following Profiler Upper: 2-Wavelength Fluorometer",
"ooi-gp03flma-ris01-05-flortd000": "Global Station Papa: Flanking Subsurface Mooring A: Mooring Riser: 3-Wavelength Fluorometer",
"ooi-gp03flmb-ris01-05-flortd000": "Global Station Papa: Flanking Subsurface Mooring B: Mooring Riser: 3-Wavelength Fluorometer",
"ooi-gi02hypm-wfp02-01-flordl000": "Global Irminger Sea: Apex Profiler Mooring: Wire-Following Profiler Upper: 2-Wavelength Fluorometer",
"ooi-gi01sumo-rii11-02-flordg033": "Global Irminger Sea: Apex Surface Mooring: Mooring Riser: 2-Wavelength Fluorometer (130 meters)",
"ooi-gi01sumo-rii11-02-flordg031": "Global Irminger Sea: Apex Surface Mooring: Mooring Riser: 2-Wavelength Fluorometer (40 meters)",
"ooi-gi01sumo-rii11-02-flordg032": "Global Irminger Sea: Apex Surface Mooring: Mooring Riser: 2-Wavelength Fluorometer (80 meters)",
"ooi-gi01sumo-rid16-02-flortd000": "Global Irminger Sea: Apex Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-gi01sumo-sbd12-02-flortd000": "Global Irminger Sea: Apex Surface Mooring: Surface Buoy: 3-Wavelength Fluorometer",
"ooi-gi03flma-ris01-05-flortd000": "Global Irminger Sea: Flanking Subsurface Mooring A: Mooring Riser: 3-Wavelength Fluorometer",
"ooi-gi03flmb-ris01-05-flortd000": "Global Irminger Sea: Flanking Subsurface Mooring B: Mooring Riser: 3-Wavelength Fluorometer",
"ooi-ga02hypm-wfp03-01-flordl000": "Global Argentine Basin: Apex Profiler Mooring: Wire-Following Profiler Lower: 2-Wavelength Fluorometer",
"ooi-ga02hypm-wfp02-01-flordl000": "Global Argentine Basin: Apex Profiler Mooring: Wire-Following Profiler Upper: 2-Wavelength Fluorometer",
"ooi-ga01sumo-rii11-02-flordg033": "Global Argentine Basin: Apex Surface Mooring: Mooring Riser: 2-Wavelength Fluorometer (130 meters)",
"ooi-ga01sumo-rii11-02-flordg031": "Global Argentine Basin: Apex Surface Mooring: Mooring Riser: 2-Wavelength Fluorometer (40 meters)",
"ooi-ga01sumo-rii11-02-flordg032": "Global Argentine Basin: Apex Surface Mooring: Mooring Riser: 2-Wavelength Fluorometer (80 meters)",
"ooi-ga01sumo-rid16-02-flortd000": "Global Argentine Basin: Apex Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-ga01sumo-sbd12-02-flortd000": "Global Argentine Basin: Apex Surface Mooring: Surface Buoy: 3-Wavelength Fluorometer",
"ooi-ga03flma-ris01-05-flortd000": "Global Argentine Basin: Flanking Subsurface Mooring A: Mooring Riser: 3-Wavelength Fluorometer",
"ooi-ga03flmb-ris01-05-flortd000": "Global Argentine Basin: Flanking Subsurface Mooring B: Mooring Riser: 3-Wavelength Fluorometer",
"ooi-gs02hypm-wfp03-01-flordl000": "Global Southern Ocean: Apex Profiler Mooring: Wire-Following Profiler Lower: 2-Wavelength Fluorometer",
"ooi-gs02hypm-wfp02-01-flordl000": "Global Southern Ocean: Apex Profiler Mooring: Wire-Following Profiler Upper: 2-Wavelength Fluorometer",
"ooi-gs01sumo-rii11-02-flordg033": "Global Southern Ocean: Apex Surface Mooring: Mooring Riser: 2-Wavelength Fluorometer (130 meters)",
"ooi-gs01sumo-rii11-02-flordg031": "Global Southern Ocean: Apex Surface Mooring: Mooring Riser: 2-Wavelength Fluorometer (40 meters)",
"ooi-gs01sumo-rii11-02-flordg032": "Global Southern Ocean: Apex Surface Mooring: Mooring Riser: 2-Wavelength Fluorometer (80 meters)",
"ooi-gs01sumo-rid16-02-flortd000": "Global Southern Ocean: Apex Surface Mooring: Near Surface Instrument Frame: 3-Wavelength Fluorometer",
"ooi-gs01sumo-sbd12-02-flortd000": "Global Southern Ocean: Apex Surface Mooring: Surface Buoy: 3-Wavelength Fluorometer",
"ooi-gs03flma-ris01-05-flortd000": "Global Southern Ocean: Flanking Subsurface Mooring A: Mooring Riser: 3-Wavelength Fluorometer",
"ooi-gs03flmb-ris01-05-flortd000": "Global Southern Ocean: Flanking Subsurface Mooring B: Mooring Riser: 3-Wavelength Fluorometer",
}
Read data from ERDDAP and save per-platform daily CHLA files#
This (optional) cell loops over all platforms in endurance_dict and:
Builds an ERDDAP URL for each dataset.
Calls
ooi_mooring_save_file(...)to:Download CHLA data.
Process it into daily 10 m-binned profiles.
Save the result as a parquet file in
_temp_data/ooi/.
The cell is wrapped in %%script false so it will not run by default;
uncomment it when you are ready to download data.
%%script false --no-raise-error
# comment to run
from pathlib import Path
out_dir = Path("_temp_data/ooi")
out_dir.mkdir(parents=True, exist_ok=True)
mooring_chl = ["ooi-ce04ossm-rid27-02-flortd000",
"ooi-cp10cnsm-rid27-02-flortd000",
"ooi-cp11nosm-rid27-02-flortd000",
"ooi-cp11sosm-rid27-02-flortd000",
"ooi-cp01cnsm-rid27-02-flortd000",
"ooi-cp03issm-rid27-02-flortd000",
"ooi-cp04ossm-rid27-02-flortd000",
"ooi-rs03axps-pc03a-4c-flordd303",
"ooi-rs01sbps-pc01a-4c-flordd103",
"ooi-gp03flma-ris01-05-flortd000",
"ooi-gp03flmb-ris01-05-flortd000",
"ooi-gi01sumo-rii11-02-flordg033",
"ooi-gi01sumo-rii11-02-flordg031",
"ooi-gi01sumo-rii11-02-flordg032",
"ooi-gi01sumo-rid16-02-flortd000",
"ooi-gi01sumo-sbd12-02-flortd000",
"ooi-gi03flma-ris01-05-flortd000",
"ooi-gi03flmb-ris01-05-flortd000",
"ooi-ga01sumo-rii11-02-flordg033",
"ooi-ga01sumo-rii11-02-flordg031",
"ooi-ga01sumo-rii11-02-flordg032",
"ooi-ga01sumo-rid16-02-flortd000",
"ooi-ga01sumo-sbd12-02-flortd000",
"ooi-ga03flma-ris01-05-flortd000",
"ooi-ga03flmb-ris01-05-flortd000",
"ooi-gs01sumo-rii11-02-flordg033",
"ooi-gs01sumo-rii11-02-flordg031",
"ooi-gs01sumo-rii11-02-flordg032",
"ooi-gs01sumo-rid16-02-flortd000",
"ooi-gs01sumo-sbd12-02-flortd000",
"ooi-gs03flma-ris01-05-flortd000",
"ooi-gs03flmb-ris01-05-flortd000",
]
for dataset_id, desc in ooi_dict.items():
# choose chlorophyll variable name
if dataset_id in mooring_chl:
chl_var = "mass_concentration_of_chlorophyll_a_in_sea_water"
else:
chl_var = "mass_concentration_of_chlorophyll_a_in_sea_water_profiler_depth_enabled"
# expected parquet path for this dataset
out_path = out_dir / f"{dataset_id}.parquet"
# skip if file already exists
if out_path.exists():
print(f"Skipping {dataset_id}: {out_path} already exists")
continue
bad = [
"ooi-cp02pmci-wfp01-04-flortk000", # no data in time range
"ooi-cp02pmco-wfp01-04-flortk000", # no data in time range
"ooi-cp01cnpm-wfp01-04-flortk000", # no data in time range
"ooi-cp01cnsm-rid27-02-flortd000", # no data in time range
"ooi-cp01cnsp-sp001-09-flortj000", # no data in time range
"ooi-cp03ispm-wfp01-04-flortk000", # no data in time range
"ooi-cp03issm-rid27-02-flortd000", # no data in time range
"ooi-cp03issp-sp001-09-flortj000", # no data in time range
"ooi-cp04ospm-wfp01-04-flortk000", # no data in time range
"ooi-cp04ossm-rid27-02-flortd000", # no data in time range
"ooi-cp02pmui-wfp01-04-flortk000", # no data in time range
"ooi-cp02pmuo-wfp01-04-flortk000", # no data in time range
"ooi-rs01sbpd-dp01a-04-flntua102", # no data in time range
"ooi-ga02hypm-wfp03-01-flordl000", # no data in time range
"ooi-ga02hypm-wfp02-01-flordl000", # no data in time range
"ooi-ga01sumo-rii11-02-flordg033", # no data in time range
"ooi-ga01sumo-rii11-02-flordg031", # no data in time range
"ooi-ga01sumo-rii11-02-flordg032", # no data in time range
"ooi-ga01sumo-rid16-02-flortd000", # no data in time range
"ooi-ga01sumo-sbd12-02-flortd000", # no data in time range
"ooi-ga03flma-ris01-05-flortd000", # no data in time range
"ooi-ga03flmb-ris01-05-flortd000", # no data in time range
"ooi-gs02hypm-wfp03-01-flordl000", # no data in time range
"ooi-gs02hypm-wfp02-01-flordl000", # no data in time range
"ooi-gs01sumo-rii11-02-flordg033", # no data in time range
"ooi-gs01sumo-rii11-02-flordg031", # no data in time range
"ooi-gs01sumo-rii11-02-flordg032", # no data in time range
"ooi-gs01sumo-rid16-02-flortd000", # no data in time range
"ooi-gs01sumo-sbd12-02-flortd000", # no data in time range
"ooi-gs03flma-ris01-05-flortd000", # no data in time range
"ooi-gs03flmb-ris01-05-flortd000", # no data in time range
"ooi-ce02shsp-sp002-07-flortj000", # no qc
"ooi-rs01sbps-pc01a-4c-flordd103", # no qc
"ooi-gp02hypm-wfp03-01-flordl000", # 2000m and deeper
]
# build URL (special case without qc_agg)
if dataset_id in bad :
continue
else:
url = (
"https://erddap.dataexplorer.oceanobservatories.org/erddap/tabledap/"
f"{dataset_id}.csv?"
"time%2Clatitude%2Clongitude%2Cz%2C"
f"{chl_var}%2C"
f"{chl_var}_qc_agg"
"&time%3E=2024-03-01"
"&time%3C=2025-12-05T00%3A00%3A00Z"
"&z%3E=-200"
)
print(f"Trying {dataset_id} with {url}")
df, saved_path = ooi_mooring_save_file(
url,
description=desc,
chl_var = chl_var,
id=dataset_id,
save=True,
out_dir=out_dir,
)
print(f"Saved {dataset_id} ({desc}) to {saved_path}")
Skipping ooi-ce04ossm-rid27-02-flortd000: _temp_data/ooi/ooi-ce04ossm-rid27-02-flortd000.parquet already exists
Skipping ooi-ce04osps-sf01b-3a-flortd104: _temp_data/ooi/ooi-ce04osps-sf01b-3a-flortd104.parquet already exists
Skipping ooi-ce09ospm-wfp01-04-flortk000: _temp_data/ooi/ooi-ce09ospm-wfp01-04-flortk000.parquet already exists
Skipping ooi-ce01issp-sp001-08-flortj000: _temp_data/ooi/ooi-ce01issp-sp001-08-flortj000.parquet already exists
Skipping ooi-ce04ospd-dp01b-04-flntua103: _temp_data/ooi/ooi-ce04ospd-dp01b-04-flntua103.parquet already exists
Skipping ooi-ce02shsp-sp001-07-flortj000: _temp_data/ooi/ooi-ce02shsp-sp001-07-flortj000.parquet already exists
Skipping ooi-ce06issp-sp001-08-flortj000: _temp_data/ooi/ooi-ce06issp-sp001-08-flortj000.parquet already exists
Skipping ooi-ce07shsp-sp001-07-flortj000: _temp_data/ooi/ooi-ce07shsp-sp001-07-flortj000.parquet already exists
Skipping ooi-cp10cnsm-rid27-02-flortd000: _temp_data/ooi/ooi-cp10cnsm-rid27-02-flortd000.parquet already exists
Skipping ooi-cp13eapm-wfp01-04-flortk000: _temp_data/ooi/ooi-cp13eapm-wfp01-04-flortk000.parquet already exists
Skipping ooi-cp14nepm-wfp01-04-flortk000: _temp_data/ooi/ooi-cp14nepm-wfp01-04-flortk000.parquet already exists
Skipping ooi-cp13nopm-wfp01-04-flortk000: _temp_data/ooi/ooi-cp13nopm-wfp01-04-flortk000.parquet already exists
Skipping ooi-cp11nosm-rid27-02-flortd000: _temp_data/ooi/ooi-cp11nosm-rid27-02-flortd000.parquet already exists
Skipping ooi-cp14sepm-wfp01-04-flortk000: _temp_data/ooi/ooi-cp14sepm-wfp01-04-flortk000.parquet already exists
Skipping ooi-cp13sopm-wfp01-04-flortk000: _temp_data/ooi/ooi-cp13sopm-wfp01-04-flortk000.parquet already exists
Skipping ooi-cp11sosm-rid27-02-flortd000: _temp_data/ooi/ooi-cp11sosm-rid27-02-flortd000.parquet already exists
Skipping ooi-rs03axps-pc03a-4c-flordd303: _temp_data/ooi/ooi-rs03axps-pc03a-4c-flordd303.parquet already exists
Skipping ooi-rs03axps-sf03a-3a-flortd301: _temp_data/ooi/ooi-rs03axps-sf03a-3a-flortd301.parquet already exists
Skipping ooi-rs01sbps-sf01a-3a-flortd101: _temp_data/ooi/ooi-rs01sbps-sf01a-3a-flortd101.parquet already exists
Skipping ooi-gp02hypm-wfp02-01-flordl000: _temp_data/ooi/ooi-gp02hypm-wfp02-01-flordl000.parquet already exists
Skipping ooi-gp03flma-ris01-05-flortd000: _temp_data/ooi/ooi-gp03flma-ris01-05-flortd000.parquet already exists
Skipping ooi-gp03flmb-ris01-05-flortd000: _temp_data/ooi/ooi-gp03flmb-ris01-05-flortd000.parquet already exists
Skipping ooi-gi02hypm-wfp02-01-flordl000: _temp_data/ooi/ooi-gi02hypm-wfp02-01-flordl000.parquet already exists
Skipping ooi-gi01sumo-rii11-02-flordg033: _temp_data/ooi/ooi-gi01sumo-rii11-02-flordg033.parquet already exists
Skipping ooi-gi01sumo-rii11-02-flordg031: _temp_data/ooi/ooi-gi01sumo-rii11-02-flordg031.parquet already exists
Skipping ooi-gi01sumo-rii11-02-flordg032: _temp_data/ooi/ooi-gi01sumo-rii11-02-flordg032.parquet already exists
Skipping ooi-gi01sumo-rid16-02-flortd000: _temp_data/ooi/ooi-gi01sumo-rid16-02-flortd000.parquet already exists
Skipping ooi-gi01sumo-sbd12-02-flortd000: _temp_data/ooi/ooi-gi01sumo-sbd12-02-flortd000.parquet already exists
Skipping ooi-gi03flma-ris01-05-flortd000: _temp_data/ooi/ooi-gi03flma-ris01-05-flortd000.parquet already exists
Skipping ooi-gi03flmb-ris01-05-flortd000: _temp_data/ooi/ooi-gi03flmb-ris01-05-flortd000.parquet already exists
Inspect a single saved daily CHLA file#
After running the ERDDAP download step, inspect one parquet file to verify:
CHLA depth-bin columns are present and look reasonable.
Time, latitude, and longitude are as expected for the platform.
# Examine the file
from pathlib import Path
import pandas as pd
out_dir = Path("_temp_data/ooi")
dataset_id = "ooi-ce01issp-sp001-08-flortj000"
out_path = f"{out_dir}/{dataset_id}.parquet"
df = pd.read_parquet(out_path)
df.head()
| time | lat | lon | CHLA_0_10 | CHLA_10_20 | CHLA_20_30 | description | |
|---|---|---|---|---|---|---|---|
| 0 | 2024-04-11 00:00:00+00:00 | 44.65845 | -124.0979 | 1.248099 | 1.369662 | 1.073702 | Coastal Endurance: Oregon Inshore Surface Pier... |
| 1 | 2024-04-12 00:00:00+00:00 | 44.65845 | -124.0979 | 1.285849 | 1.286291 | 0.988821 | Coastal Endurance: Oregon Inshore Surface Pier... |
| 2 | 2024-04-17 00:00:00+00:00 | 44.65845 | -124.0979 | 0.384099 | 0.510783 | 0.457698 | Coastal Endurance: Oregon Inshore Surface Pier... |
| 3 | 2024-04-18 00:00:00+00:00 | 44.65845 | -124.0979 | 0.330928 | 0.311356 | 0.278780 | Coastal Endurance: Oregon Inshore Surface Pier... |
| 4 | 2024-04-19 00:00:00+00:00 | 44.65845 | -124.0979 | 0.384770 | 0.366793 | 0.327334 | Coastal Endurance: Oregon Inshore Surface Pier... |
Merge all platforms into a single OOI CHLA dataset#
Here we:
Read all per-platform parquet files from
_temp_data/ooi/.Ensure that each file has all CHLA depth-bin columns (
CHLA_0_10, …,CHLA_190_200), filling missing bins withNaN.Add a
profile_idof the formdatasetID_YYYYMMDDso each daily profile is uniquely identified by platform × day.Concatenate all platforms into a single dataframe and sort rows by time.
Save the merged dataset to
_temp_data/ooi_chla_0_200m.parquet.
#%%script false --no-raise-error
import numpy as np
import pandas as pd
from pathlib import Path
# directory where individual mooring parquet files live
out_dir = Path("_temp_data/ooi")
# all the CHLA columns you want to guarantee
chla_bins = [f"CHLA_{b}_{b+10}" for b in range(0, 200, 10)]
dfs = []
for p in out_dir.glob("*.parquet"):
df = pd.read_parquet(p)
# dataset_id from filename, e.g. "ooi-ce04osps-sf01b-3a-flortd104"
dataset_id = p.stem
# ensure time is datetime and make a date column
df["time"] = pd.to_datetime(df["time"])
date = df["time"].dt.floor("D")
# profile_id = dataset_id_YYYYMMDD
df["profile_id"] = dataset_id + "_" + date.dt.strftime("%Y%m%d")
# ensure all CHLA_* columns exist
for col in chla_bins:
if col not in df.columns:
df[col] = np.nan
# consistent column order
base_cols = ["profile_id", "time", "lat", "lon"]
all_chla_cols = chla_bins
other_cols = [c for c in df.columns if c not in base_cols + all_chla_cols]
df = df[base_cols + all_chla_cols + other_cols]
dfs.append(df)
# combine all rows
combined = pd.concat(dfs, ignore_index=True)
# sort by time (date) across all platforms
combined["time"] = pd.to_datetime(combined["time"])
combined = combined.sort_values("time").reset_index(drop=True)
# optional: save
combined_out = Path("_temp_data/ooi_chla_0_200m.parquet")
combined.to_parquet(combined_out)
combined.head()
| profile_id | time | lat | lon | CHLA_0_10 | CHLA_10_20 | CHLA_20_30 | CHLA_30_40 | CHLA_40_50 | CHLA_50_60 | ... | CHLA_110_120 | CHLA_120_130 | CHLA_130_140 | CHLA_140_150 | CHLA_150_160 | CHLA_160_170 | CHLA_170_180 | CHLA_180_190 | CHLA_190_200 | description | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | ooi-ce04ossm-rid27-02-flortd000_20240301 | 2024-03-01 00:00:00+00:00 | 44.365280 | -124.939470 | 0.675962 | NaN | NaN | NaN | NaN | NaN | ... | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | Coastal Endurance: Oregon Offshore Surface Moo... |
| 1 | ooi-gi03flmb-ris01-05-flortd000_20240301 | 2024-03-01 00:00:00+00:00 | 59.715220 | -39.315150 | 0.348521 | NaN | NaN | NaN | NaN | NaN | ... | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | Global Irminger Sea: Flanking Subsurface Moori... |
| 2 | ooi-rs03axps-sf03a-3a-flortd301_20240301 | 2024-03-01 00:00:00+00:00 | 45.830466 | -129.753082 | NaN | 1.032399 | 1.128244 | 1.216156 | 1.239003 | 1.200323 | ... | 0.031526 | 0.025329 | 0.020974 | 0.020147 | 0.020719 | 0.018667 | 0.023159 | 0.026216 | NaN | Regional Cabled Array: Axial Base Shallow Prof... |
| 3 | ooi-ce09ospm-wfp01-04-flortk000_20240301 | 2024-03-01 00:00:00+00:00 | 46.852180 | -124.974610 | NaN | NaN | NaN | 0.712121 | 0.697653 | 0.647610 | ... | 0.029593 | 0.025464 | 0.023151 | 0.022770 | 0.021980 | 0.023111 | 0.022429 | 0.022202 | 0.024052 | Coastal Endurance: Washington Offshore Profile... |
| 4 | ooi-gp02hypm-wfp02-01-flordl000_20240301 | 2024-03-01 00:00:00+00:00 | 50.070370 | -144.800750 | NaN | NaN | NaN | NaN | NaN | NaN | ... | NaN | NaN | NaN | NaN | NaN | 0.030767 | 0.037275 | 0.038340 | 0.031950 | Global Station Papa: Apex Profiler Mooring: Wi... |
5 rows × 25 columns
Select a random daily profile and plot CHLA vs depth#
Finally, we:
Load the merged Endurance CHLA dataset.
Identify
profile_idvalues with at least a minimum number of non-NaN CHLA depth bins.Randomly select one valid daily profile.
Retrieve the platform description for context.
Convert CHLA depth-bin columns into depth midpoints and plot CHLA vs depth for the selected daily profile.
# Create a plot
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from pathlib import Path
import textwrap
# ---- 1. Load combined dataset ----
combined_path = Path("_temp_data/ooi_chla_0_200m.parquet")
df = pd.read_parquet(combined_path)
# Ensure time is datetime (not strictly needed for the plot, but nice to have)
df["time"] = pd.to_datetime(df["time"])
# ---- 2. Identify CHLA columns and pick a good profile ----
chla_cols = [c for c in df.columns if c.startswith("CHLA_")]
if not chla_cols:
raise ValueError("No CHLA_ columns found in the dataframe.")
# Count how many non-NaN CHLA values each profile_id has (per row)
# (Assumes one row per profile_id; if multiple, takes the max across rows.)
counts = (
df.groupby("profile_id")[chla_cols]
.apply(lambda x: x.notna().sum(axis=1).max())
)
# Get a profile_id with at least 10 valid CHLA bins
valid_profiles = counts[counts >= 10]
if valid_profiles.empty:
raise ValueError("No profile_id found with at least 10 non-NaN CHLA bins.")
rand_row = valid_profiles.sample(1, random_state=None) # set an int seed if you want reproducible
profile_id = rand_row.index[0]
n_bins = rand_row.iloc[0]
print(f"Using profile_id: {profile_id} with {n_bins} valid CHLA bins")
# Get Description
desc = df.loc[df["profile_id"] == profile_id, "description"].iloc[0]
# ---- 3. Extract that profile ----
sub = df[df["profile_id"] == profile_id].copy()
# If there somehow are multiple rows for this profile, average them
if len(sub) > 1:
sub = sub.groupby("profile_id")[chla_cols].mean().reset_index()
else:
sub = sub.reset_index(drop=True)
# Take the first (or only) row of CHLA values
chla_vals = sub[chla_cols].iloc[0].values
# ---- 4. Build depth midpoints from CHLA_0_10 style column names ----
# CHLA_0_10 -> lower edge = 0 -> midpoint = 5
depths = []
for col in chla_cols:
# col format assumed: "CHLA_<z1>_<z2>"
parts = col.split("_")
if len(parts) < 3:
raise ValueError(f"Unexpected CHLA column format: {col}")
z1 = int(parts[1])
z_mid = z1 + 5 # 10 m bin midpoint
depths.append(z_mid)
depths = np.array(depths)
chla_vals = np.array(chla_vals, dtype=float)
# Sort by depth (just in case columns are out of order)
order = np.argsort(depths)
depths = depths[order]
chla_vals = chla_vals[order]
# Mask out NaNs so the plot doesn't connect them
valid = ~np.isnan(chla_vals)
depths_valid = depths[valid]
chla_valid = chla_vals[valid]
# ---- 5. Plot CHLA vs depth ----
plt.figure(figsize=(6, 8))
plt.plot(chla_valid, depths_valid, marker="o")
plt.gca().invert_yaxis() # surface at top
plt.xlabel("CHLA (mg m$^{-3}$)")
plt.ylabel("Depth (m)")
wrapped_desc = textwrap.fill(desc, width=50) # adjust width to taste
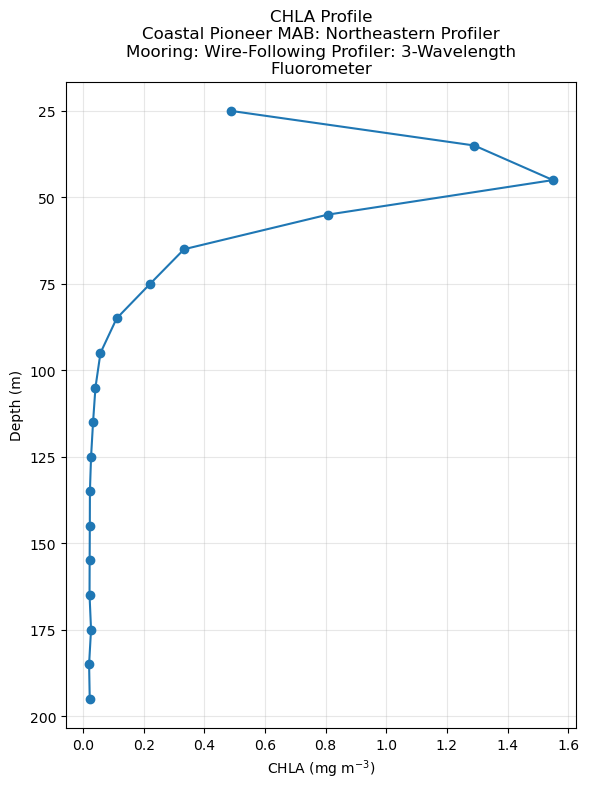
plt.title(f"CHLA Profile\n{wrapped_desc}")
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
Using profile_id: ooi-cp14nepm-wfp01-04-flortk000_20240629 with 18 valid CHLA bins
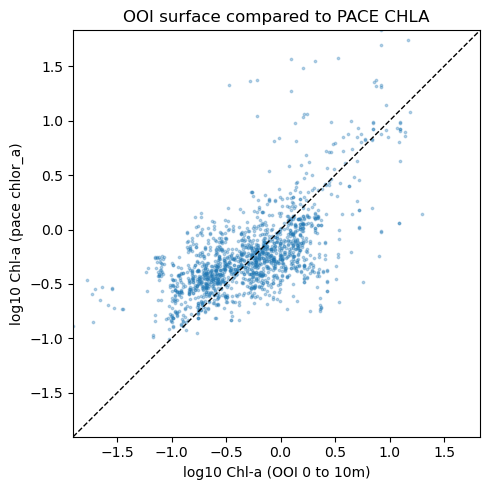
Match ups to PACE#
# --- Custom python functions ---
import os, importlib
# Looks to see if you have the file already and if not, downloads from GitHub
if not os.path.exists("ml_utils.py"):
!wget -q https://raw.githubusercontent.com/fish-pace/2025-tutorials/main/ml_utils.py
import ml_utils as mu
importlib.reload(mu)
<module 'ml_utils' from '/home/jovyan/fish-pace-datasets/ml_utils.py'>
# Load data from GitHub
df_y_name="CHLA"; df_lat_name="lat"; df_lon_name="lon"; df_time_name="time"
import pandas as pd
base_url="https://raw.githubusercontent.com/fish-pace/fish-pace-datasets/main/datasets/chla_z/data"
url = f"{base_url}/CHLA_ooi_profiles.parquet"
ooi_chl_df = pd.read_parquet(url)
len(ooi_chl_df)
9782
%%time
from datetime import datetime
def run_batch(
results, df,
ds_vec_name="wavelength", ds_var_name="Rrs", ds_vec_sel=None,
df_time_name="time", df_lat_name="lat", df_lon_name="lon"
):
"""
Run a batch of PACE files (results) and return PACE variables for lat/lon/time rows in a
dataframe.
Parameters
----------
results : earthaccess results object as returned by `earthaccess.search`).
df : A pandas DataFrame containing Argo observations. Must include columns for
time, latitude, longitude.
ds_vec_name : str or None, optional
Name of the spectral dimension in the PACE dataset (e.g. "wavelength").
If not None, matched satellite spectra are returned with one column per
wavelength. If None, only a single variable is extracted.
ds_vec_sel : value or None, optional
Value of the spectral dimension in the PACE dataset (e.g. "wavelength")
to select. If None, matched satellite spectra are returned with one column per
wavelength. If given, only a single variable is extracted for that value.
ds_var_name : str, optional
Name of the variable to extract from the PACE dataset
(e.g. "Rrs" or "chlor_a").
df_lat_name, df_lon_name, df_time_name : str, optional
Column names in `df` for latitude, longitude, and time.
"""
# Make sure to refresh fileset to minimize the chance that the token expires before we are done
# this for loop consumes about 6Gb of RAM
fileset = earthaccess.open(results, pqdm_kwargs={"disable": True} );
df_plus = []
for i, f in enumerate(fileset):
df_record, pts = mu.one_file_matches(
f, df,
ds_vec_name=ds_vec_name, ds_var_name=ds_var_name, ds_vec_sel=ds_vec_sel,
df_time_name=df_time_name, df_lat_name=df_lat_name, df_lon_name=df_lon_name)
if df_record is None or len(df_record) == 0:
print(f"Skipped day {i} no data in df")
continue
# error check
if len(df_record) != len(pts):
raise ValueError(f"Row mismatch: df_record={len(df_record)}, pts={len(pts)}")
# left "concat": keep df_record rows, add pts columns by position
df_record_plus = pd.concat(
[
df_record.reset_index(drop=True),
pts.reset_index(drop=True),
], axis=1,)
df_plus.append(df_record_plus)
if not len(df_plus) == 0:
df_plus = pd.concat(df_plus, ignore_index=True)
return df_plus
CPU times: user 13 μs, sys: 1 μs, total: 14 μs
Wall time: 17.6 μs
# Load data from GitHub
df_y_name="CHLA"; df_lat_name="LATITUDE"; df_lon_name="LONGITUDE"; df_time_name="TIME"
import pandas as pd
base_url="https://raw.githubusercontent.com/fish-pace/2025-tutorials/main/data/argo_monthly_nc/"
url = f"{base_url}argo_profiles_index.parquet"
argo_chl_df = pd.read_parquet(url)
len(argo_chl_df)
Get Rrs#
import earthaccess
auth = earthaccess.login()
# are we authenticated?
if not auth.authenticated:
# ask for credentials and persist them in a .netrc file
auth.login(strategy="interactive", persist=True)
import xarray as xr
df = ooi_chl_df
rrs_results = earthaccess.search_data(
short_name = "PACE_OCI_L3M_RRS",
temporal = (df.time.min(), df.time.max()),
granule_name="*.DAY.*.4km.*"
)
rrs_results_nrt = earthaccess.search_data(
short_name = "PACE_OCI_L3M_RRS_NRT",
temporal = (df.time.min(), df.time.max()),
granule_name="*.DAY.*.4km.*"
)
results = rrs_results + rrs_results_nrt
print(f"{len(results)} days of PACE data")
718 days of PACE data
from pathlib import Path
from datetime import datetime
import pandas as pd
results_dir = Path("_temp_data/matchups/ooi")
results_dir.mkdir(exist_ok=True)
all_batches = []
batch_size = 10
var = "CHLA"
results = results
df = ooi_chl_df
ds_var = "Rrs"
for batch_idx, i in enumerate(range(0, len(results), batch_size), start=1):
batch = results[i:i+batch_size]
print(f"Batch {batch_idx}: {len(batch)} files: {datetime.now():%H:%M:%S}")
# If this batch was already done, skip it
batch_path = results_dir / f"{var}_matchups_{ds_var}_batch_{batch_idx:03d}.parquet"
if batch_path.exists():
print(f" -> Skipping batch {batch_idx}, found {batch_path}")
continue
df_batch = run_batch(batch, df) # returns a DataFrame
# Save this batch immediately
df_batch.to_parquet(batch_path, index=False)
print(f" -> Saved {len(df_batch)} rows to {batch_path}")
Batch 1: 10 files: 18:57:44
-> Saved 118 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_001.parquet
Batch 2: 10 files: 18:58:08
-> Saved 109 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_002.parquet
Batch 3: 10 files: 18:58:24
-> Saved 109 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_003.parquet
Batch 4: 10 files: 18:58:44
-> Saved 168 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_004.parquet
Batch 5: 10 files: 18:59:13
-> Saved 190 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_005.parquet
Batch 6: 10 files: 18:59:45
-> Saved 180 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_006.parquet
Batch 7: 10 files: 19:00:15
-> Saved 183 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_007.parquet
Batch 8: 10 files: 19:00:47
-> Saved 189 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_008.parquet
Batch 9: 10 files: 19:01:19
-> Saved 185 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_009.parquet
Batch 10: 10 files: 19:01:50
-> Saved 222 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_010.parquet
Batch 11: 10 files: 19:02:25
-> Saved 203 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_011.parquet
Batch 12: 10 files: 19:03:01
-> Saved 189 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_012.parquet
Batch 13: 10 files: 19:03:34
-> Saved 188 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_013.parquet
Batch 14: 10 files: 19:04:08
-> Saved 175 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_014.parquet
Batch 15: 10 files: 19:04:40
-> Saved 156 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_015.parquet
Batch 16: 10 files: 19:05:11
-> Saved 152 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_016.parquet
Batch 17: 10 files: 19:05:42
-> Saved 173 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_017.parquet
Batch 18: 10 files: 19:06:15
-> Saved 213 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_018.parquet
Batch 19: 10 files: 19:06:51
-> Saved 189 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_019.parquet
Batch 20: 10 files: 19:07:24
-> Saved 170 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_020.parquet
Batch 21: 10 files: 19:07:53
-> Saved 163 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_021.parquet
Batch 22: 10 files: 19:08:22
-> Saved 177 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_022.parquet
Batch 23: 10 files: 19:08:49
-> Saved 188 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_023.parquet
Batch 24: 10 files: 19:09:16
-> Saved 167 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_024.parquet
Batch 25: 10 files: 19:09:42
-> Saved 169 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_025.parquet
Batch 26: 10 files: 19:10:06
-> Saved 163 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_026.parquet
Batch 27: 10 files: 19:10:31
-> Saved 153 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_027.parquet
Batch 28: 10 files: 19:10:51
-> Saved 142 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_028.parquet
Batch 29: 10 files: 19:11:12
-> Saved 146 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_029.parquet
Batch 30: 10 files: 19:11:32
-> Saved 141 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_030.parquet
Batch 31: 10 files: 19:11:52
-> Saved 145 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_031.parquet
Batch 32: 10 files: 19:12:14
-> Saved 144 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_032.parquet
Batch 33: 10 files: 19:12:35
-> Saved 138 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_033.parquet
Batch 34: 10 files: 19:12:57
-> Saved 129 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_034.parquet
Batch 35: 10 files: 19:13:18
-> Saved 126 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_035.parquet
Batch 36: 10 files: 19:13:40
-> Saved 112 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_036.parquet
Batch 37: 10 files: 19:14:00
-> Saved 100 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_037.parquet
Batch 38: 10 files: 19:14:20
-> Saved 97 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_038.parquet
Batch 39: 10 files: 19:14:38
-> Saved 94 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_039.parquet
Batch 40: 10 files: 19:14:56
-> Saved 133 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_040.parquet
Batch 41: 10 files: 19:15:18
-> Saved 135 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_041.parquet
Batch 42: 10 files: 19:15:43
-> Saved 123 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_042.parquet
Batch 43: 10 files: 19:16:05
-> Saved 129 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_043.parquet
Batch 44: 10 files: 19:16:30
-> Saved 161 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_044.parquet
Batch 45: 10 files: 19:16:59
-> Saved 168 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_045.parquet
Batch 46: 10 files: 19:17:26
-> Saved 155 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_046.parquet
Batch 47: 10 files: 19:17:53
-> Saved 157 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_047.parquet
Batch 48: 10 files: 19:18:23
-> Saved 149 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_048.parquet
Batch 49: 10 files: 19:18:54
-> Saved 155 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_049.parquet
Batch 50: 10 files: 19:19:24
-> Saved 174 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_050.parquet
Batch 51: 10 files: 19:19:57
-> Saved 169 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_051.parquet
Batch 52: 10 files: 19:20:30
-> Saved 173 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_052.parquet
Batch 53: 10 files: 19:21:04
-> Saved 172 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_053.parquet
Batch 54: 10 files: 19:21:37
-> Saved 170 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_054.parquet
Batch 55: 10 files: 19:22:10
-> Saved 158 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_055.parquet
Batch 56: 10 files: 19:22:41
-> Saved 146 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_056.parquet
Batch 57: 10 files: 19:23:12
-> Saved 152 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_057.parquet
Batch 58: 10 files: 19:23:42
-> Saved 146 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_058.parquet
Batch 59: 10 files: 19:24:14
-> Saved 176 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_059.parquet
Batch 60: 10 files: 19:24:45
-> Saved 169 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_060.parquet
Batch 61: 10 files: 19:25:17
-> Saved 174 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_061.parquet
Batch 62: 10 files: 19:25:51
-> Saved 174 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_062.parquet
Batch 63: 10 files: 19:26:24
-> Saved 173 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_063.parquet
Batch 64: 10 files: 19:26:57
-> Saved 159 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_064.parquet
Batch 65: 10 files: 19:27:27
-> Saved 153 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_065.parquet
Batch 66: 10 files: 19:27:58
-> Saved 122 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_066.parquet
Batch 67: 10 files: 19:28:23
-> Saved 130 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_067.parquet
Batch 68: 10 files: 19:28:47
-> Saved 130 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_068.parquet
Batch 69: 10 files: 19:29:10
-> Saved 117 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_069.parquet
Batch 70: 10 files: 19:29:30
-> Saved 92 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_070.parquet
Batch 71: 10 files: 19:29:46
-> Saved 90 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_071.parquet
Batch 72: 8 files: 19:30:03
Skipped day 7 no data in df
-> Saved 50 rows to _temp_data/matchups/ooi/CHLA_matchups_Rrs_batch_072.parquet
# When everything is done, you can merge all batches:
from pathlib import Path
import pandas as pd
results_dir = Path("_temp_data/matchups/ooi")
var = "CHLA"
ds_var = "Rrs"
batch_files = sorted(results_dir.glob(f"{var}_matchups_{ds_var}_batch_*.parquet"))
dfs = [pd.read_parquet(f) for f in batch_files]
df_all = pd.concat(dfs, ignore_index=True)
df_all.to_parquet(f"{results_dir}/{var}_ooi_{ds_var}_all.parquet", index=False)
print(f"Total rows: {len(df_all)}")
Total rows: 10989
Get chlor_a#
# Add on chla to the clean parquet
import pandas as pd
import earthaccess
import xarray as xr
time_range = (ooi_chl_df.time.min(), ooi_chl_df.time.max())
# Get PACE CHL data
import earthaccess
auth = earthaccess.login()
# are we authenticated?
if not auth.authenticated:
# ask for credentials and persist them in a .netrc file
auth.login(strategy="interactive", persist=True)
import xarray as xr
chl_results = earthaccess.search_data(
short_name = "PACE_OCI_L3M_CHL",
temporal = time_range,
granule_name="*.DAY.*.4km.*"
)
chl_results_nrt = earthaccess.search_data(
short_name = "PACE_OCI_L3M_CHL_NRT",
temporal = time_range,
granule_name="*.DAY.*.4km.*"
)
results = chl_results + chl_results_nrt
print(f"{len(results)} days of PACE data")
718 days of PACE data
from pathlib import Path
from datetime import datetime
import pandas as pd
results_dir = Path("_temp_data/matchups/ooi")
results_dir.mkdir(exist_ok=True)
all_batches = []
batch_size = 10
var = "CHLA"
results = results
df = ooi_chl_df
ds_var = "chlor_a"
for batch_idx, i in enumerate(range(0, len(results), batch_size), start=1):
batch = results[i:i+batch_size]
print(f"Batch {batch_idx}: {len(batch)} files: {datetime.now():%H:%M:%S}")
# If this batch was already done, skip it
batch_path = results_dir / f"{var}_matchups_{ds_var}_batch_{batch_idx:03d}.parquet"
if batch_path.exists():
print(f" -> Skipping batch {batch_idx}, found {batch_path}")
continue
df_batch = run_batch(batch, df, ds_vec_name=None, ds_var_name="chlor_a")
# Save this batch immediately
df_batch.to_parquet(batch_path, index=False)
print(f" -> Saved {len(df_batch)} rows to {batch_path}")
Batch 1: 10 files: 20:04:20
-> Saved 118 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_001.parquet
Batch 2: 10 files: 20:04:30
-> Saved 109 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_002.parquet
Batch 3: 10 files: 20:04:35
-> Saved 109 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_003.parquet
Batch 4: 10 files: 20:04:39
-> Saved 168 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_004.parquet
Batch 5: 10 files: 20:04:44
-> Saved 190 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_005.parquet
Batch 6: 10 files: 20:04:49
-> Saved 180 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_006.parquet
Batch 7: 10 files: 20:04:54
-> Saved 183 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_007.parquet
Batch 8: 10 files: 20:05:00
-> Saved 189 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_008.parquet
Batch 9: 10 files: 20:05:05
-> Saved 185 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_009.parquet
Batch 10: 10 files: 20:05:10
-> Saved 222 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_010.parquet
Batch 11: 10 files: 20:05:16
-> Saved 203 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_011.parquet
Batch 12: 10 files: 20:05:21
-> Saved 189 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_012.parquet
Batch 13: 10 files: 20:05:26
-> Saved 188 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_013.parquet
Batch 14: 10 files: 20:05:31
-> Saved 175 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_014.parquet
Batch 15: 10 files: 20:05:36
-> Saved 156 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_015.parquet
Batch 16: 10 files: 20:05:40
-> Saved 152 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_016.parquet
Batch 17: 10 files: 20:05:45
-> Saved 173 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_017.parquet
Batch 18: 10 files: 20:05:49
-> Saved 213 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_018.parquet
Batch 19: 10 files: 20:05:55
-> Saved 189 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_019.parquet
Batch 20: 10 files: 20:06:00
-> Saved 170 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_020.parquet
Batch 21: 10 files: 20:06:04
-> Saved 163 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_021.parquet
Batch 22: 10 files: 20:06:09
-> Saved 177 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_022.parquet
Batch 23: 10 files: 20:06:13
-> Saved 188 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_023.parquet
Batch 24: 10 files: 20:06:18
-> Saved 167 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_024.parquet
Batch 25: 10 files: 20:06:23
-> Saved 169 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_025.parquet
Batch 26: 10 files: 20:06:27
-> Saved 163 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_026.parquet
Batch 27: 10 files: 20:06:32
-> Saved 153 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_027.parquet
Batch 28: 10 files: 20:06:36
-> Saved 142 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_028.parquet
Batch 29: 10 files: 20:06:40
-> Saved 146 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_029.parquet
Batch 30: 10 files: 20:06:45
-> Saved 141 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_030.parquet
Batch 31: 10 files: 20:06:48
-> Saved 145 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_031.parquet
Batch 32: 10 files: 20:06:52
-> Saved 144 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_032.parquet
Batch 33: 10 files: 20:06:57
-> Saved 138 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_033.parquet
Batch 34: 10 files: 20:07:01
-> Saved 129 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_034.parquet
Batch 35: 10 files: 20:07:05
-> Saved 126 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_035.parquet
Batch 36: 10 files: 20:07:09
-> Saved 112 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_036.parquet
Batch 37: 10 files: 20:07:13
-> Saved 100 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_037.parquet
Batch 38: 10 files: 20:07:18
-> Saved 97 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_038.parquet
Batch 39: 10 files: 20:07:21
-> Saved 94 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_039.parquet
Batch 40: 10 files: 20:07:25
-> Saved 133 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_040.parquet
Batch 41: 10 files: 20:07:29
-> Saved 135 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_041.parquet
Batch 42: 10 files: 20:07:33
-> Saved 123 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_042.parquet
Batch 43: 10 files: 20:07:37
-> Saved 129 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_043.parquet
Batch 44: 10 files: 20:07:42
-> Saved 161 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_044.parquet
Batch 45: 10 files: 20:07:46
-> Saved 168 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_045.parquet
Batch 46: 10 files: 20:07:50
-> Saved 155 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_046.parquet
Batch 47: 10 files: 20:07:55
-> Saved 157 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_047.parquet
Batch 48: 10 files: 20:07:59
-> Saved 149 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_048.parquet
Batch 49: 10 files: 20:08:04
-> Saved 155 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_049.parquet
Batch 50: 10 files: 20:08:08
-> Saved 174 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_050.parquet
Batch 51: 10 files: 20:08:12
-> Saved 169 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_051.parquet
Batch 52: 10 files: 20:08:17
-> Saved 173 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_052.parquet
Batch 53: 10 files: 20:08:22
-> Saved 172 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_053.parquet
Batch 54: 10 files: 20:08:26
-> Saved 170 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_054.parquet
Batch 55: 10 files: 20:08:31
-> Saved 158 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_055.parquet
Batch 56: 10 files: 20:08:35
-> Saved 146 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_056.parquet
Batch 57: 10 files: 20:08:40
-> Saved 152 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_057.parquet
Batch 58: 10 files: 20:08:44
-> Saved 146 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_058.parquet
Batch 59: 10 files: 20:08:49
-> Saved 176 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_059.parquet
Batch 60: 10 files: 20:08:54
-> Saved 169 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_060.parquet
Batch 61: 10 files: 20:08:58
-> Saved 174 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_061.parquet
Batch 62: 10 files: 20:09:04
-> Saved 174 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_062.parquet
Batch 63: 10 files: 20:09:08
-> Saved 173 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_063.parquet
Batch 64: 10 files: 20:09:13
-> Saved 159 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_064.parquet
Batch 65: 10 files: 20:09:17
-> Saved 153 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_065.parquet
Batch 66: 10 files: 20:09:22
-> Saved 122 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_066.parquet
Batch 67: 10 files: 20:09:26
-> Saved 130 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_067.parquet
Batch 68: 10 files: 20:09:31
-> Saved 130 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_068.parquet
Batch 69: 10 files: 20:09:35
-> Saved 117 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_069.parquet
Batch 70: 10 files: 20:09:39
-> Saved 92 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_070.parquet
Batch 71: 10 files: 20:09:43
-> Saved 90 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_071.parquet
Batch 72: 8 files: 20:09:46
Skipped day 7 no data in df
-> Saved 50 rows to _temp_data/matchups/ooi/CHLA_matchups_chlor_a_batch_072.parquet
# When everything is done, you can merge all batches:
from pathlib import Path
import pandas as pd
results_dir = Path("_temp_data/matchups/ooi")
platform = "ooi"
var = "CHLA"
ds_var = "chlor_a"
batch_files = sorted(results_dir.glob(f"{var}_matchups_{ds_var}_batch_*.parquet"))
dfs = [pd.read_parquet(f) for f in batch_files]
df_all = pd.concat(dfs, ignore_index=True)
df_all.to_parquet(f"{results_dir}/{var}_{platform}_{ds_var}_all.parquet", index=False)
Kd_490#
# Add on chla to the clean parquet
import pandas as pd
import earthaccess
import xarray as xr
time_range = (ooi_chl_df.time.min(), ooi_chl_df.time.max())
# Get PACE CHL data
import earthaccess
auth = earthaccess.login()
# are we authenticated?
if not auth.authenticated:
# ask for credentials and persist them in a .netrc file
auth.login(strategy="interactive", persist=True)
import xarray as xr
kd_results = earthaccess.search_data(
short_name = "PACE_OCI_L3M_KD",
temporal = time_range,
granule_name="*.DAY.*.4km.*"
)
kd_results_nrt = earthaccess.search_data(
short_name = "PACE_OCI_L3M_KD_NRT",
temporal = time_range,
granule_name="*.DAY.*.4km.*"
)
results = kd_results + kd_results_nrt
print(f"{len(results)} days of PACE data")
718 days of PACE data
from pathlib import Path
from datetime import datetime
import pandas as pd
results_dir = Path("_temp_data/matchups/ooi")
results_dir.mkdir(exist_ok=True)
all_batches = []
batch_size = 10
var = "CHLA"
results = results
df = ooi_chl_df
ds_var = "Kd"
for batch_idx, i in enumerate(range(0, len(results), batch_size), start=1):
batch = results[i:i+batch_size]
print(f"Batch {batch_idx}: {len(batch)} files: {datetime.now():%H:%M:%S}")
# If this batch was already done, skip it
batch_path = results_dir / f"{var}_matchups_{ds_var}_batch_{batch_idx:03d}.parquet"
if batch_path.exists():
print(f" -> Skipping batch {batch_idx}, found {batch_path}")
continue
df_batch = run_batch(batch, ooi_chl_df, ds_var_name="Kd", ds_vec_sel=490.0)
# Save this batch immediately
df_batch.to_parquet(batch_path, index=False)
print(f" -> Saved {len(df_batch)} rows to {batch_path}")
Batch 1: 10 files: 20:24:00
-> Saved 118 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_001.parquet
Batch 2: 10 files: 20:24:24
-> Saved 109 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_002.parquet
Batch 3: 10 files: 20:24:30
-> Saved 109 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_003.parquet
Batch 4: 10 files: 20:24:35
-> Saved 168 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_004.parquet
Batch 5: 10 files: 20:24:42
-> Saved 190 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_005.parquet
Batch 6: 10 files: 20:24:49
-> Saved 180 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_006.parquet
Batch 7: 10 files: 20:24:56
-> Saved 183 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_007.parquet
Batch 8: 10 files: 20:25:03
-> Saved 189 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_008.parquet
Batch 9: 10 files: 20:25:10
-> Saved 185 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_009.parquet
Batch 10: 10 files: 20:25:16
-> Saved 222 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_010.parquet
Batch 11: 10 files: 20:25:24
-> Saved 203 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_011.parquet
Batch 12: 10 files: 20:25:32
-> Saved 189 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_012.parquet
Batch 13: 10 files: 20:25:39
-> Saved 188 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_013.parquet
Batch 14: 10 files: 20:25:46
-> Saved 175 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_014.parquet
Batch 15: 10 files: 20:25:54
-> Saved 156 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_015.parquet
Batch 16: 10 files: 20:26:02
-> Saved 152 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_016.parquet
Batch 17: 10 files: 20:26:09
-> Saved 173 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_017.parquet
Batch 18: 10 files: 20:26:17
-> Saved 213 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_018.parquet
Batch 19: 10 files: 20:26:24
-> Saved 189 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_019.parquet
Batch 20: 10 files: 20:26:32
-> Saved 170 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_020.parquet
Batch 21: 10 files: 20:26:40
-> Saved 163 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_021.parquet
Batch 22: 10 files: 20:26:48
-> Saved 177 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_022.parquet
Batch 23: 10 files: 20:26:54
-> Saved 188 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_023.parquet
Batch 24: 10 files: 20:27:01
-> Saved 167 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_024.parquet
Batch 25: 10 files: 20:27:06
-> Saved 169 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_025.parquet
Batch 26: 10 files: 20:27:12
-> Saved 163 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_026.parquet
Batch 27: 10 files: 20:27:18
-> Saved 153 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_027.parquet
Batch 28: 10 files: 20:27:23
-> Saved 142 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_028.parquet
Batch 29: 10 files: 20:27:28
-> Saved 146 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_029.parquet
Batch 30: 10 files: 20:27:33
-> Saved 141 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_030.parquet
Batch 31: 10 files: 20:27:38
-> Saved 145 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_031.parquet
Batch 32: 10 files: 20:27:43
-> Saved 144 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_032.parquet
Batch 33: 10 files: 20:27:49
-> Saved 138 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_033.parquet
Batch 34: 10 files: 20:27:54
-> Saved 129 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_034.parquet
Batch 35: 10 files: 20:27:58
-> Saved 126 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_035.parquet
Batch 36: 10 files: 20:28:04
-> Saved 112 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_036.parquet
Batch 37: 10 files: 20:28:09
-> Saved 100 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_037.parquet
Batch 38: 10 files: 20:28:15
-> Saved 97 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_038.parquet
Batch 39: 10 files: 20:28:20
-> Saved 94 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_039.parquet
Batch 40: 10 files: 20:28:25
-> Saved 133 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_040.parquet
Batch 41: 10 files: 20:28:31
-> Saved 135 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_041.parquet
Batch 42: 10 files: 20:28:37
-> Saved 123 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_042.parquet
Batch 43: 10 files: 20:28:43
-> Saved 129 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_043.parquet
Batch 44: 10 files: 20:28:49
-> Saved 161 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_044.parquet
Batch 45: 10 files: 20:28:55
-> Saved 168 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_045.parquet
Batch 46: 10 files: 20:29:01
-> Saved 155 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_046.parquet
Batch 47: 10 files: 20:29:07
-> Saved 157 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_047.parquet
Batch 48: 10 files: 20:29:13
-> Saved 149 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_048.parquet
Batch 49: 10 files: 20:29:21
-> Saved 155 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_049.parquet
Batch 50: 10 files: 20:29:28
-> Saved 174 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_050.parquet
Batch 51: 10 files: 20:29:35
-> Saved 169 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_051.parquet
Batch 52: 10 files: 20:29:43
-> Saved 173 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_052.parquet
Batch 53: 10 files: 20:29:50
-> Saved 172 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_053.parquet
Batch 54: 10 files: 20:29:58
-> Saved 170 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_054.parquet
Batch 55: 10 files: 20:30:06
-> Saved 158 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_055.parquet
Batch 56: 10 files: 20:30:13
-> Saved 146 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_056.parquet
Batch 57: 10 files: 20:30:21
-> Saved 152 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_057.parquet
Batch 58: 10 files: 20:30:28
-> Saved 146 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_058.parquet
Batch 59: 10 files: 20:30:35
-> Saved 176 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_059.parquet
Batch 60: 10 files: 20:30:43
-> Saved 169 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_060.parquet
Batch 61: 10 files: 20:30:50
-> Saved 174 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_061.parquet
Batch 62: 10 files: 20:30:58
-> Saved 174 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_062.parquet
Batch 63: 10 files: 20:31:05
-> Saved 173 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_063.parquet
Batch 64: 10 files: 20:31:13
-> Saved 159 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_064.parquet
Batch 65: 10 files: 20:31:20
-> Saved 153 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_065.parquet
Batch 66: 10 files: 20:31:28
-> Saved 122 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_066.parquet
Batch 67: 10 files: 20:31:34
-> Saved 130 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_067.parquet
Batch 68: 10 files: 20:31:40
-> Saved 130 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_068.parquet
Batch 69: 10 files: 20:31:46
-> Saved 117 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_069.parquet
Batch 70: 10 files: 20:31:51
-> Saved 92 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_070.parquet
Batch 71: 10 files: 20:31:55
-> Saved 90 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_071.parquet
Batch 72: 8 files: 20:32:00
Skipped day 7 no data in df
-> Saved 50 rows to _temp_data/matchups/ooi/CHLA_matchups_Kd_batch_072.parquet
# When everything is done, you can merge all batches:
from pathlib import Path
import pandas as pd
results_dir = Path("_temp_data/matchups/ooi")
platform = "ooi"
var = "CHLA"
ds_var = "Kd"
batch_files = sorted(results_dir.glob(f"{var}_matchups_{ds_var}_batch_*.parquet"))
dfs = [pd.read_parquet(f) for f in batch_files]
df_all = pd.concat(dfs, ignore_index=True)
df_all.to_parquet(f"{results_dir}/{var}_{platform}_{ds_var}_all.parquet", index=False)
from pathlib import Path
import pandas as pd
import numpy as np
results_dir = Path("_temp_data/matchups/ooi")
platform = "ooi"
var = "CHLA"
df_all_kd = pd.read_parquet(f"{results_dir}/{var}_{platform}_Kd_all.parquet")
df_all_chl = pd.read_parquet(f"{results_dir}/{var}_{platform}_chlor_a_all.parquet")
df_all_rrs = pd.read_parquet(f"{results_dir}/{var}_{platform}_Rrs_all.parquet")
print(df_all_kd.shape)
print(df_all_rrs.shape)
(10989, 31)
(10989, 202)
df_all_chl.columns
Index(['profile_id', 'time', 'lat', 'lon', 'CHLA_0_10', 'CHLA_10_20',
'CHLA_20_30', 'CHLA_30_40', 'CHLA_40_50', 'CHLA_50_60', 'CHLA_60_70',
'CHLA_70_80', 'CHLA_80_90', 'CHLA_90_100', 'CHLA_100_110',
'CHLA_110_120', 'CHLA_120_130', 'CHLA_130_140', 'CHLA_140_150',
'CHLA_150_160', 'CHLA_160_170', 'CHLA_170_180', 'CHLA_180_190',
'CHLA_190_200', 'description', 'pace_chlor_a_file',
'pace_chlor_a_t_start', 'pace_chlor_a_t_end', 'pace_chlor_a_lat',
'pace_chlor_a_lon', 'pace_chlor_a'],
dtype='object')
n = 5
assert np.array_equal(
df_all_chl.iloc[:, n].values,
df_all_chl.iloc[:, n].values,
equal_nan=True,
), "First n column values differ"
Merge PACE matchups all together#
# When everything is done, you can merge all batches:
from pathlib import Path
import pandas as pd
import numpy as np
results_dir = Path("_temp_data/matchups/ooi")
platform = "ooi"
var = "CHLA"
df_all_kd = pd.read_parquet(f"{results_dir}/{var}_{platform}_Kd_all.parquet")
df_all_chl = pd.read_parquet(f"{results_dir}/{var}_{platform}_chlor_a_all.parquet")
df_all_rrs = pd.read_parquet(f"{results_dir}/{var}_{platform}_Rrs_all.parquet")
# check
n = 4
assert np.array_equal(
df_all_chl.iloc[:, :n].values,
df_all_rrs.iloc[:, :n].values,
), "First n column values differ"
assert np.array_equal(
df_all_kd.iloc[:, :n].values,
df_all_rrs.iloc[:, :n].values,
), "First n column values differ"
n1 = 5
n2 = 23
assert np.array_equal(
df_all_chl.iloc[:, n1:n2].values,
df_all_rrs.iloc[:, n1:n2].values,
equal_nan=True,
), "First n column values differ"
assert np.array_equal(
df_all_kd.iloc[:, n1:n2].values,
df_all_rrs.iloc[:, n1:n2].values,
equal_nan=True,
), "First n column values differ"
# columns to add
df_chl_extra = df_all_chl["pace_chlor_a"]
df_kd_extra = df_all_kd["pace_Kd_490"]
# Horizontal concat: rrs on the left, extras on the right
df_merged = pd.concat([df_all_rrs, df_chl_extra, df_kd_extra], axis=1)
df_merged.to_parquet(f"{results_dir}/{var}_{platform}_Rrs_chlor_a_Kd_all.parquet", index=False)
Remove duplicates#
# Cleaned;
# Duplicates replaced with the PACE data that is closest to TIME
import pandas as pd
from pathlib import Path
results_dir = Path("_temp_data/matchups/ooi")
platform = "ooi"
var = "CHLA"
df_merged = pd.read_parquet(f"{results_dir}/{var}_{platform}_Rrs_chlor_a_Kd_all.parquet")
df = df_merged
# Parse all times as UTC-aware datetimes
for col in ["time", "pace_Rrs_t_start", "pace_Rrs_t_end"]:
df[col] = pd.to_datetime(df[col], utc=True)
# is file NRT
df["nrt"] = [("NRT" in f) for f in df["pace_Rrs_file"]]
idx = df.groupby("profile_id")["nrt"].idxmin()
df = df.loc[idx].drop(columns=["nrt"]).reset_index(drop=True)
# Just in case there are 2 files due to slightly overlapping time windows
# though turns out there were not any
# center of the PACE time window
df["pace_center"] = df["pace_Rrs_t_start"] + (
df["pace_Rrs_t_end"] - df["pace_Rrs_t_start"]
) / 2
# 2. absolute time difference between profile TIME and window center
df["time_diff"] = (df["time"] - df["pace_center"]).abs()
# 3. for each profile_id, keep the row with smallest time_diff
idx = df.groupby("profile_id")["time_diff"].idxmin()
df_dedup = df.loc[idx].drop(columns=["pace_center", "time_diff"]).reset_index(drop=True)
df_dedup = df_dedup.sort_values(by="time").reset_index(drop=True)
print(df_dedup.shape)
# double-check
df_dedup['profile_id'].duplicated().any()
(6415, 204)
np.False_
out_path = "data/CHLA_ooi_profiles_plus_PACE.parquet"
df_merged.to_parquet(out_path)
Create metadata#
import pyarrow as pa
import pyarrow.parquet as pq
from datetime import datetime
import pandas as pd
out_path = "data/CHLA_ooi_profiles_plus_PACE.parquet"
df = pd.read_parquet(out_path)
table = pa.Table.from_pandas(df)
file_meta = {
"title": "Global OOI CHLA profile metrics (0 to 200 m, 10 m bins) with PACE Rrs, chlor_a and Kd_490 matchups",
"creator": "Eli Holmes / NOAA https://orcid.org/0000-0001-9128-8393",
"created": datetime.utcnow().isoformat() + "Z",
"source": (
"OOI data (via ERDDAP) and Daily 4km PACE data v 3.1 collections "
"PACE_OCI_L3M_RRS, PACE_OCI_L3M_CHL and PACE_OCI_L3M_KD via NASA Earthdata. "
"NSF Ocean Observatories Initiative. (2014). Data from 2024-03-01 to 2025-12-05. Data Explorer ERDDAP. https://erddap.dataexplorer.oceanobservatories.org] doi:10.58046/OOI-CP03ISSM. Accessed on 2025-12-05."
"See profile_id column for exact platform used."
),
"description": (
"All OOI Profiler and Mooring Florometer data Mar 2024 to Nov 2025 with CHLA variable was downloaded. "
"Data accessed on https://erddap.dataexplorer.oceanobservatories.org/erddap. data_id is the profile_id in file."
"Per-profile depth-binned CHLA means (0–200 m by 10 m bins) were computed for each depth bin. "
"QC filtering was done on the values using the _qc_agg variable. Values filtered to qc=1."
"All profiles were kept even if some binned averages were missing. "
"PACE Rrs for all wavelengths, chlor_a, and Kd_490 were extracted from Level 3 4km v 3.1 Daily files. "
"Matching was done by first selecting all profiles with TIME within 24 hours of t_start in the "
"PACE Level 3 file. Note: t_start in the netcdf attributes was used, not the day in the netcdf filename or "
"in the NASA Earthdata CMS. Once all profiles within that timerange were found, the LATITUDE/LONGITUDE point "
"for the Argo profile was matched to the PACE Level 3 4km grid via ds.sel(lon=LONGITUDE, lat=LATITUDE, method='nearest') "
"where ds is an xarray Dataset formed by reading in the PACE file. "
"The actual PACE files used are listed in the dataset under pace_var_file. "
"Note: the PACE data is heavily cloud-affected and the majority (≈80%) of PACE pixels are NaN."
"Data were accessed using earthaccess Python package, example "
"results = earthaccess.search_data(short_name = 'PACE_OCI_L3M_KD', temporal = ('2024-03-01', '2025-11-30'), granule_name='*.DAY.*.4km.*')"
),
"profile_id_definition": "ERDDAP data id",
"time_definition": "time in UTC reduced to day. All samples within the same day are given the same time stamp.",
"lat_definition": "lat in the ERDDAP dataset. The OOI platforms are fixed (non-moving in lat/lon)",
"lon_definition": "lon in the ERDDAP dataset.",
"CHLA_A_B_definition": (
"Depth-binned averages of CHLA. Computed as the average of all individual CHLA measurements within the depth interval and within a day."
"(z>=A and z<B), where z is depth below surface. "
"Data were filtered to qc_agg = 1 values."
),
"pace_Rrs_lat_definition": "The matched latitude from the PACE Level 3 grid.",
"pace_Rrs_lon_definition": "The matched longitude from the PACE Level 3 grid.",
"pace_Rrs_t_start_definition": "Start time of the swath data used to create the Level 3 4km file. In UTC, from netcdf attributes.",
"pace_Rrs_t_end_definition": "End time of the swath data used to create the Level 3 4km file. In UTC, from netcdf attributes.",
"pace_Rrs_file_definition": "Filename for the Level 3 4km file. Only Rrs is shown but chlor_a and Kd_490 are analogous.",
"pace_Rrs_A_definition": (
"PACE_OCI_L3M_RRS Level 3 4km remote-sensing reflectance for wavelength A, "
"where A is the center wavelength in nanometers (e.g., pace_Rrs_443 for 443 nm), "
"extracted from PACE v 3.1 Daily files."
),
"pace_chlor_a_definition": "PACE_OCI_L3M_CHL Level 3 4km chlor_a extracted from PACE v 3.1 Daily files.",
"pace_Kd_490_definition": "PACE_OCI_L3M_KD Level 3 4km Kd_490 extracted from PACE v 3.1 Daily files.",
"variable_lat_standard_name": "latitude",
"variable_lat_units": "degrees_north",
"variable_lon_standard_name": "longitude",
"variable_lon_units": "degrees_east",
"variable_pace_Rrs_lat_standard_name": "latitude",
"variable_pace_Rrs_lat_units": "degrees_north",
"variable_pace_Rrs_lon_standard_name": "longitude",
"variable_pace_Rrs_lon_units": "degrees_east",
"variable_pace_Rrs_t_start_standard_name": "time",
"variable_pace_Rrs_t_start_units": "UTC",
"variable_pace_Rrs_t_end_standard_name": "time",
"variable_pace_Rrs_t_end_units": "UTC",
"variable_pace_CHLA_prefix_definition": (
"All columns whose names start with 'CHLA_' followed by two numbers "
"are the averaged CHLA within a depth bin defined by the numbers. "
"For example, 'CHLA_10_20' is the average CHLA between z>=10 and z<20 m."
),
"CHLA_processing_description": (
"CHLA values were taken directly from the OOI chl variable which was 'mass_concentration_of_chlorophyll_a_in_sea_water'"
"for moorings and 'mass_concentration_of_chlorophyll_a_in_sea_water_profiler_depth_enabled' for profilers."
"(mg m-3). No additional sensor corrections or non-photochemical "
"quenching adjustments were applied. QC was given in qc_agg column."
"Values were filtered to qc_agg = 1. CHLA measurements were aggregated into 10 m "
"depth bins between 0 and 200 m using arithmetic means for all values in a day."
),
"CHLA_measurement_description": (
"OOI chlorophyll-a (CHLA) is measured with WET Labs ECO fluorometers "
"(e.g., FLORT/FLORD family) deployed on Ocean Observatories Initiative "
"platforms. These sensors emit blue light (~470 nm) to excite "
"chlorophyll pigments in the water and detect the resulting "
"fluorescence in the red band near ~695 nm. Raw fluorescence counts are "
"converted to chlorophyll-a concentration using factory calibration "
"coefficients and OOI processing, which applies standard scale factors, "
"dark corrections, and automated quality-control tests to produce the "
"'fluorometric_chlor_a' data product (reported in approximately "
"mg m⁻³). In this work, we use the OOI fluorometric chlorophyll-a "
"values as provided by the OOI data system, without additional "
"post-processing (e.g., non-photochemical quenching corrections or "
"region-specific bias adjustments), so they should be interpreted as "
"instrument-based estimates of relative chlorophyll-a concentration "
"rather than fully validated absolute values."
),
"variable_CHLA_standard_name": "mass_concentration_of_chlorophyll_a_in_sea_water",
"variable_CHLA_units": "mg m-3",
"variable_pace_Rrs_prefix_definition": (
"All columns whose names start with 'pace_Rrs_' contain PACE OCI "
"remote-sensing reflectance (Rrs) at the wavelength given by the suffix "
"in nanometers. For example, 'pace_Rrs_443' is Rrs at 443 nm. "
"Rrs is the ratio of water-leaving radiance to downwelling irradiance "
"just above the sea surface."
),
"variable_pace_Rrs_long_name": "Remote sensing reflectance",
"variable_pace_Rrs_standard_name": (
"surface_ratio_of_upwelling_radiance_emerging_from_sea_water_to_"
"downwelling_radiative_flux_in_air"
),
"variable_pace_Rrs_units": "sr-1",
"Rrs_measurement_definition": (
"Remote-sensing reflectance (Rrs) is the ratio of water-leaving radiance "
"to downwelling irradiance just above the sea surface (sr-1). "
"Rrs(λ) from PACE OCI is derived from top-of-atmosphere radiances "
"after atmospheric correction that removes molecular scattering, "
"aerosol scattering, and surface glint contributions. "
"Rrs represents the spectral signature of the ocean water column "
"and is the primary input for bio-optical algorithms. "
"PACE OCI provides hyperspectral Rrs from 350–890 nm at 5 nm resolution. "
"pace_Rrs_λ variables in this dataset correspond to center wavelengths "
"equal to the value in the variable name (e.g., pace_Rrs_443 for 443 nm)."
),
"pace_chlor_a_long_name": "Chlorophyll Concentration, OCI Algorithm",
"pace_chlor_a_standard_name": "mass_concentration_of_chlorophyll_a_in_sea_water",
"pace_chlor_a_units": "mg m-3",
"pace_chlor_a_valid_min": "0.001",
"pace_chlor_a_valid_max": "100.0",
"pace_chlor_a_measurement_definition": (
"From PACE v 3.1 Level 3 files: PACE OCI chlorophyll-a concentration (chlor_a) "
"is computed using the OCI Chlorophyll Algorithm (CI algorithm) described by "
"Hu, Lee, and Franz (2012). The CI algorithm estimates chlorophyll-a "
"concentration from hyperspectral remote-sensing reflectance (Rrs) using a "
"three-band baseline difference approach optimized for clear to moderately "
"productive waters. Values represent near-surface chlorophyll-a concentration "
"in mg m-3. See: Hu, C., Lee Z., and Franz, B.A. (2012). "
"'Chlorophyll-a algorithms for oligotrophic oceans: A novel approach based on "
"three-band reflectance difference,' J. Geophys. Res., 117, C01011."
),
"pace_Kd_490_standard_name": "attenuation_of_downwelling_radiative_flux_in_sea_water",
"pace_Kd_490_units": "m-1",
"pace_Kd_490_measurement_definition": (
"Diffuse attenuation coefficient for downwelling irradiance at 490 nm (Kd_490), "
"retrieved from PACE OCI Rrs. "
"Kd_490 describes the rate at which light at 490 nm decays with depth due to "
"absorption and scattering in the water column. Higher Kd_490 values indicate "
"optically darker, more turbid, or more absorbing waters. "
"From PACE v 3.1 Level 3 files; key references include: "
"Lee, Z.P., M. Darecki, K.L. Carder, C.O. Davis, D. Stramski, W.J. Rhea, "
"'Diffuse Attenuation coefficient of downwelling irradiance: An evaluation of remote sensing methods' (2005), "
"doi:10.1029/2004JC002573; and Zhongping Lee, Chuanmin Hu, Shaoling Shang, "
"Keping Du, Marlon Lewis, Robert Arnone, and Robert Brewin, "
"'Penetration of UV-visible solar radiation in the global oceans: Insights from ocean color remote sensing' (2013), "
"doi:10.1002/jgrc.20308."
),
"spatiotemporal_coverage_time_start": "2024-03-01T00:00:00Z",
"spatiotemporal_coverage_time_end": "2025-12-05T23:59:59Z",
"spatiotemporal_coverage_lat_min": "-90.0",
"spatiotemporal_coverage_lat_max": "90.0",
"spatiotemporal_coverage_lon_min": "-180.0",
"spatiotemporal_coverage_lon_max": "180.0",
"license": "Open access; unrestricted use with attribution."
}
table = table.replace_schema_metadata(file_meta)
out_path = "data/CHLA_ooi_profiles_plus_PACE.parquet"
pq.write_table(table, out_path)
# Check metadata
import pyarrow.parquet as pq
out_path = "data/CHLA_ooi_profiles_plus_PACE.parquet"
t = pq.read_table(out_path)
t.schema.metadata
{b'title': b'Global OOI CHLA profile metrics (0 to 200 m, 10 m bins) with PACE Rrs, chlor_a and Kd_490 matchups',
b'creator': b'Eli Holmes / NOAA https://orcid.org/0000-0001-9128-8393',
b'created': b'2025-12-08T21:52:26.092100Z',
b'source': b'OOI data (via ERDDAP) and Daily 4km PACE data v 3.1 collections PACE_OCI_L3M_RRS, PACE_OCI_L3M_CHL and PACE_OCI_L3M_KD via NASA Earthdata. NSF Ocean Observatories Initiative. (2014). Data from 2024-03-01 to 2025-12-05. Data Explorer ERDDAP. https://erddap.dataexplorer.oceanobservatories.org] doi:10.58046/OOI-CP03ISSM. Accessed on 2025-12-05.See profile_id column for exact platform used.',
b'description': b"All OOI Profiler and Mooring Florometer data Mar 2024 to Nov 2025 with CHLA variable was downloaded. Data accessed on https://erddap.dataexplorer.oceanobservatories.org/erddap. data_id is the profile_id in file.Per-profile depth-binned CHLA means (0\xe2\x80\x93200 m by 10 m bins) were computed for each depth bin. QC filtering was done on the values using the _qc_agg variable. Values filtered to qc=1.All profiles were kept even if some binned averages were missing. PACE Rrs for all wavelengths, chlor_a, and Kd_490 were extracted from Level 3 4km v 3.1 Daily files. Matching was done by first selecting all profiles with TIME within 24 hours of t_start in the PACE Level 3 file. Note: t_start in the netcdf attributes was used, not the day in the netcdf filename or in the NASA Earthdata CMS. Once all profiles within that timerange were found, the LATITUDE/LONGITUDE point for the Argo profile was matched to the PACE Level 3 4km grid via ds.sel(lon=LONGITUDE, lat=LATITUDE, method='nearest') where ds is an xarray Dataset formed by reading in the PACE file. The actual PACE files used are listed in the dataset under pace_var_file. Note: the PACE data is heavily cloud-affected and the majority (\xe2\x89\x8880%) of PACE pixels are NaN.Data were accessed using earthaccess Python package, example results = earthaccess.search_data(short_name = 'PACE_OCI_L3M_KD', temporal = ('2024-03-01', '2025-11-30'), granule_name='*.DAY.*.4km.*')",
b'profile_id_definition': b'ERDDAP data id',
b'time_definition': b'time in UTC reduced to day. All samples within the same day are given the same time stamp.',
b'lat_definition': b'lat in the ERDDAP dataset. The OOI platforms are fixed (non-moving in lat/lon)',
b'lon_definition': b'lon in the ERDDAP dataset.',
b'CHLA_A_B_definition': b'Depth-binned averages of CHLA. Computed as the average of all individual CHLA measurements within the depth interval and within a day.(z>=A and z<B), where z is depth below surface. Data were filtered to qc_agg = 1 values.',
b'pace_Rrs_lat_definition': b'The matched latitude from the PACE Level 3 grid.',
b'pace_Rrs_lon_definition': b'The matched longitude from the PACE Level 3 grid.',
b'pace_Rrs_t_start_definition': b'Start time of the swath data used to create the Level 3 4km file. In UTC, from netcdf attributes.',
b'pace_Rrs_t_end_definition': b'End time of the swath data used to create the Level 3 4km file. In UTC, from netcdf attributes.',
b'pace_Rrs_file_definition': b'Filename for the Level 3 4km file. Only Rrs is shown but chlor_a and Kd_490 are analogous.',
b'pace_Rrs_A_definition': b'PACE_OCI_L3M_RRS Level 3 4km remote-sensing reflectance for wavelength A, where A is the center wavelength in nanometers (e.g., pace_Rrs_443 for 443 nm), extracted from PACE v 3.1 Daily files.',
b'pace_chlor_a_definition': b'PACE_OCI_L3M_CHL Level 3 4km chlor_a extracted from PACE v 3.1 Daily files.',
b'pace_Kd_490_definition': b'PACE_OCI_L3M_KD Level 3 4km Kd_490 extracted from PACE v 3.1 Daily files.',
b'variable_lat_standard_name': b'latitude',
b'variable_lat_units': b'degrees_north',
b'variable_lon_standard_name': b'longitude',
b'variable_lon_units': b'degrees_east',
b'variable_pace_Rrs_lat_standard_name': b'latitude',
b'variable_pace_Rrs_lat_units': b'degrees_north',
b'variable_pace_Rrs_lon_standard_name': b'longitude',
b'variable_pace_Rrs_lon_units': b'degrees_east',
b'variable_pace_Rrs_t_start_standard_name': b'time',
b'variable_pace_Rrs_t_start_units': b'UTC',
b'variable_pace_Rrs_t_end_standard_name': b'time',
b'variable_pace_Rrs_t_end_units': b'UTC',
b'variable_pace_CHLA_prefix_definition': b"All columns whose names start with 'CHLA_' followed by two numbers are the averaged CHLA within a depth bin defined by the numbers. For example, 'CHLA_10_20' is the average CHLA between z>=10 and z<20 m.",
b'CHLA_processing_description': b"CHLA values were taken directly from the OOI chl variable which was 'mass_concentration_of_chlorophyll_a_in_sea_water'for moorings and 'mass_concentration_of_chlorophyll_a_in_sea_water_profiler_depth_enabled' for profilers.(mg m-3). No additional sensor corrections or non-photochemical quenching adjustments were applied. QC was given in qc_agg column.Values were filtered to qc_agg = 1. CHLA measurements were aggregated into 10 m depth bins between 0 and 200 m using arithmetic means for all values in a day.",
b'CHLA_measurement_description': b"OOI chlorophyll-a (CHLA) is measured with WET Labs ECO fluorometers (e.g., FLORT/FLORD family) deployed on Ocean Observatories Initiative platforms. These sensors emit blue light (~470 nm) to excite chlorophyll pigments in the water and detect the resulting fluorescence in the red band near ~695 nm. Raw fluorescence counts are converted to chlorophyll-a concentration using factory calibration coefficients and OOI processing, which applies standard scale factors, dark corrections, and automated quality-control tests to produce the 'fluorometric_chlor_a' data product (reported in approximately mg m\xe2\x81\xbb\xc2\xb3). In this work, we use the OOI fluorometric chlorophyll-a values as provided by the OOI data system, without additional post-processing (e.g., non-photochemical quenching corrections or region-specific bias adjustments), so they should be interpreted as instrument-based estimates of relative chlorophyll-a concentration rather than fully validated absolute values.",
b'variable_CHLA_standard_name': b'mass_concentration_of_chlorophyll_a_in_sea_water',
b'variable_CHLA_units': b'mg m-3',
b'variable_pace_Rrs_prefix_definition': b"All columns whose names start with 'pace_Rrs_' contain PACE OCI remote-sensing reflectance (Rrs) at the wavelength given by the suffix in nanometers. For example, 'pace_Rrs_443' is Rrs at 443 nm. Rrs is the ratio of water-leaving radiance to downwelling irradiance just above the sea surface.",
b'variable_pace_Rrs_long_name': b'Remote sensing reflectance',
b'variable_pace_Rrs_standard_name': b'surface_ratio_of_upwelling_radiance_emerging_from_sea_water_to_downwelling_radiative_flux_in_air',
b'variable_pace_Rrs_units': b'sr-1',
b'Rrs_measurement_definition': b'Remote-sensing reflectance (Rrs) is the ratio of water-leaving radiance to downwelling irradiance just above the sea surface (sr-1). Rrs(\xce\xbb) from PACE OCI is derived from top-of-atmosphere radiances after atmospheric correction that removes molecular scattering, aerosol scattering, and surface glint contributions. Rrs represents the spectral signature of the ocean water column and is the primary input for bio-optical algorithms. PACE OCI provides hyperspectral Rrs from 350\xe2\x80\x93890 nm at 5 nm resolution. pace_Rrs_\xce\xbb variables in this dataset correspond to center wavelengths equal to the value in the variable name (e.g., pace_Rrs_443 for 443 nm).',
b'pace_chlor_a_long_name': b'Chlorophyll Concentration, OCI Algorithm',
b'pace_chlor_a_standard_name': b'mass_concentration_of_chlorophyll_a_in_sea_water',
b'pace_chlor_a_units': b'mg m-3',
b'pace_chlor_a_valid_min': b'0.001',
b'pace_chlor_a_valid_max': b'100.0',
b'pace_chlor_a_measurement_definition': b"From PACE v 3.1 Level 3 files: PACE OCI chlorophyll-a concentration (chlor_a) is computed using the OCI Chlorophyll Algorithm (CI algorithm) described by Hu, Lee, and Franz (2012). The CI algorithm estimates chlorophyll-a concentration from hyperspectral remote-sensing reflectance (Rrs) using a three-band baseline difference approach optimized for clear to moderately productive waters. Values represent near-surface chlorophyll-a concentration in mg m-3. See: Hu, C., Lee Z., and Franz, B.A. (2012). 'Chlorophyll-a algorithms for oligotrophic oceans: A novel approach based on three-band reflectance difference,' J. Geophys. Res., 117, C01011.",
b'pace_Kd_490_standard_name': b'attenuation_of_downwelling_radiative_flux_in_sea_water',
b'pace_Kd_490_units': b'm-1',
b'pace_Kd_490_measurement_definition': b"Diffuse attenuation coefficient for downwelling irradiance at 490 nm (Kd_490), retrieved from PACE OCI Rrs. Kd_490 describes the rate at which light at 490 nm decays with depth due to absorption and scattering in the water column. Higher Kd_490 values indicate optically darker, more turbid, or more absorbing waters. From PACE v 3.1 Level 3 files; key references include: Lee, Z.P., M. Darecki, K.L. Carder, C.O. Davis, D. Stramski, W.J. Rhea, 'Diffuse Attenuation coefficient of downwelling irradiance: An evaluation of remote sensing methods' (2005), doi:10.1029/2004JC002573; and Zhongping Lee, Chuanmin Hu, Shaoling Shang, Keping Du, Marlon Lewis, Robert Arnone, and Robert Brewin, 'Penetration of UV-visible solar radiation in the global oceans: Insights from ocean color remote sensing' (2013), doi:10.1002/jgrc.20308.",
b'spatiotemporal_coverage_time_start': b'2024-03-01T00:00:00Z',
b'spatiotemporal_coverage_time_end': b'2025-12-05T23:59:59Z',
b'spatiotemporal_coverage_lat_min': b'-90.0',
b'spatiotemporal_coverage_lat_max': b'90.0',
b'spatiotemporal_coverage_lon_min': b'-180.0',
b'spatiotemporal_coverage_lon_max': b'180.0',
b'license': b'Open access; unrestricted use with attribution.'}
# Create or update the STAC entry for the PACE matchup file
collection_path = "data/data_collection.json"
collection = mu.load_or_create_collection(collection_path)
pace_item_id = "global-ooi-chla-profile-metrics-0-200m-10m-bins-with-pace-oci-matchups"
pace_file_name = "CHLA_ooi_profiles_plus_PACE.parquet"
# raw GitHub URL pattern
pace_href = f"https://raw.githubusercontent.com/fish-pace/fish-pace-datasets/main/data/{pace_file_name}"
# Notebook that creates / documents this dataset
pace_notebook_href = "https://github.com/fish-pace/fish-pace-datasets/blob/main/ooi.ipynb"
collection = mu.add_or_update_item(
collection,
item_id=pace_item_id,
asset_href=pace_href,
title=(
"Global OOI CHLA profile metrics (0–200 m, 10 m bins) "
"with PACE Rrs, chlor_a, and Kd_490 matchups"
),
description=(
"Depth-binned CHLA metrics from all OOI Profilers and Moorings from Mar 2024 to Dec 2025, "
"Depth-binned from 0–200 m in 10 m bins using unadjusted CHLA values, "
"matched to PACE OCI Level-3 4 km v3.1 Daily Rrs, chlor_a, and Kd_490. "
"Profiles are matched to PACE Level-3 pixels within 24 hours of the file t_start "
"attribute and to the nearest 4 km grid cell in latitude/longitude."
),
start_datetime="2024-03-01T00:00:00Z",
end_datetime="2025-12-05T23:59:59Z",
extra_properties={
"license": "Open access; unrestricted use with attribution.",
"variables": ["CHLA", "Rrs", "chlor_a", "Kd_490"],
"platform": "OOI Profilers and Moorings + PACE OCI",
"tutorial_notebook": pace_notebook_href,
"file_name": pace_file_name
}
)
mu.save_collection(collection, collection_path)
mu.stac_to_readme(
"data/data_collection.json",
readme_path="data/README.md",
repo_raw_base="https://raw.githubusercontent.com/fish-pace/fish-pace-datasets/main"
)
README.md written to data/README.md
Summary#
This completes the preparation of our OOI + PACE matchups dataset. Now that we have the data we can use is to train a model to predict CHLA using OOI CHLA data and compare to PACE chlor_a from Rrs ratios.
The OOI and PACE (satellite-derived) chlorophyll-a estimates will not match. One is based on reflectance from the water column while the other is in-situ reflectance. But they are correlated. This plot shows a oft observed pattern where the satellite derived estimate is higher when surface CHLA is lower. Conversely, the in-situ surface measurement is a bit higher when CHLA is higher. But note that we don’t have many in-situ measurements over 1 (on log10 scale) so about 10 on the non-log scale. According to the PACE measurements we do have values above 1.4 (bloom), but the OOI measurements tend to be lower in those cases.
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import r2_score, mean_squared_error
df = pd.read_parquet("data/CHLA_ooi_profiles_plus_PACE.parquet")
df_clean = df.dropna(subset=["CHLA_0_10"])
# Take logs (natural log; use np.log10 if you prefer log10)
y_ooi = df_clean["CHLA_0_10"].values
y_pace = df_clean["pace_chlor_a"].values
# Mask: positive & finite in both
mask = (
np.isfinite(y_ooi) &
np.isfinite(y_pace) &
(y_ooi > 0) &
(y_pace > 0)
)
y_ooi = y_ooi[mask]
y_pace = y_pace[mask]
y_ooi = np.log10(y_ooi)
y_pace = np.log10(y_pace)
# Scatter plot
plt.figure(figsize=(5, 5))
plt.scatter(y_ooi, y_pace, s=3, alpha=0.3)
# 1:1 line
lims = [
min(y_ooi.min(), y_pace.min()),
max(y_ooi.max(), y_pace.max())
]
plt.plot(lims, lims, "k--", linewidth=1)
plt.xlim(lims)
plt.ylim(lims)
plt.xlabel("log10 Chl-a (OOI 0 to 10m)")
plt.ylabel("log10 Chl-a (pace chlor_a)")
plt.title("OOI surface compared to PACE CHLA")
plt.tight_layout()
plt.show()